CLADES: a programmable sequence of reporters for lineage analysis
Preprint posted on 30 May 2019 https://www.biorxiv.org/content/10.1101/655308v1
CLADES - An innovative strategy to resolve lineage progression in an organism through multicolor labeling.
Selected by Ying-Tsen TungCategories: developmental biology, genetics, molecular biology
Background:
Lineage tracing identifies all the descendants generated from a single ancestor cell, which provides important information about how cell fates are progressively acquired during tissue development. Several revolutionary techniques have been developed for lineage tracing [1]. Imaging-based systems utilize genetic approaches to induce the expression of randomly combined fluorescent reporters in different lineages. These methods reveal cell clones in situ but cannot trace lineage progression. On the other hand, DNA sequencing based strategies rely on the CRISPR/Cas9 system to stochastically generate DNA barcodes that progressively accumulate over cell divisions, therefore allowing to deduce the birth order of lineages at large-scale. However, both spatial and morphological information is lost with this approach. In this preprint, the authors developed CLADES (Cell Lineage Access Driven by an Edition Sequence) as a programmable imaging based tool to trace cell clones and resolve their lineage progression by pre-established color order.
Key findings:
CLADES utilizes programmable genetic switches to induce or inactivate fluorescent reporters sequentially. The combination of different reporters generates a pre-determined color code to reveal the birth order of cells in a lineage.
The genetic switch builds upon a gRNA cascade where each gRNA is sequentially activated via DNA editing mediated by its corresponding pre-activated gRNA. There are two keys. One is to make the editing outcome predictable, and the other is to express the gRNA structure in an inactive form and make it functional after the predictable DNA editing. For this purpose, the authors utilized CaSSA, a CRISPR based approach they developed in a recent preprint [2] (highlighted by a preLight post [3]), which is designed to allow single strand annealing (SSA) to mediate precise recombination upon Cas9 editing. Here, the authors optimized a gRNA structure with a CaSSA switch in the scaffold region to achieve minimal leakiness without the trigger gRNA in Drosophila and zebrafish.
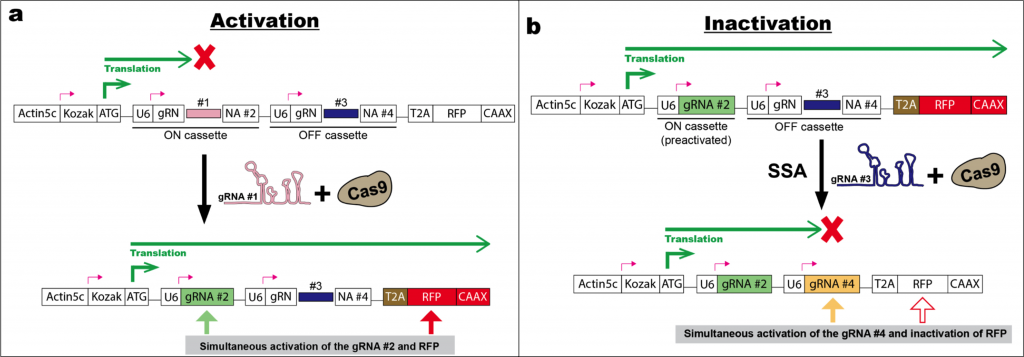
Another critical step of the CLADES design is to synchronize the cascade of gRNAs and reporters. Ideally, preactivated gRNA#1 directs Cas9 to simultaneously activate the reporter#1 (ex. GFP) and gRNA#2, which in turn switches on the reporter #2 (ex.RFP). Yet the authors found a failure of reporter progression when the reporter and gRNA loci were independently positioned in Drosophila. After several rounds of optimization, they alternatively embed gRNA cassettes within the targeted reporter which is actively expressed as a truncated non-functional protein. The reporter contains two gRNA switching cassettes, one for making the reporter in-frame upon editing (ON cassette) and one for placing the same reporter out-of-frame upon editing (OFF cassette) (Fig. 1). This approach ensures the sequential regulation of reporters by corresponding gRNA and maximizes the lineage tracing capacity.
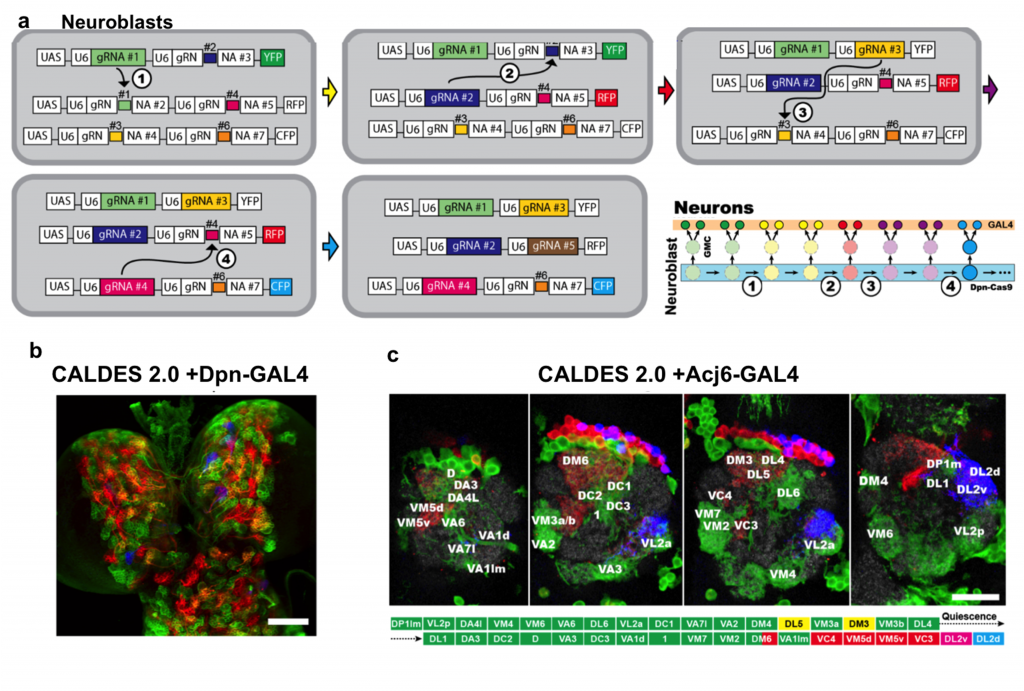
Based on the elegant design, the authors developed the advanced CLADES 2.0 system with 3 reporters and 5 colors in Drosophila. It combined a serial ON and OFF cassettes to generate gRNA cascades for the sequential expression of YFP → YFP+RFP → RFP → RFP+CPF → CFP. The authors also incorporated the GAL4/UAS system to drive reporters that only label lineages with specific cell identities. When testing CLADES 2.0 in all neuroblasts, the larva brain can be labeled by all five predicted colors. Furthermore, the progression of the 5 color cascade can also reflect the birth order in the well-characterized ALad1 lineage (Fig. 2).
In addition to lineage tracing, CLADES has great potential to reveal any sequential biological event. By using a germline-specific driver for Cas9 expression in CLADES 1.0, the color of the dominant fly population can refer generations.
It should be noted that CLADES is still at a suboptimal status and can be further improved. The progression efficiency gradually reduced over each step, mainly due to the occurrence of non-SSA mediated recombination events (~40%). Therefore, a portion of the progeny loses the tracers, especially for the late-born cells. Also, the occasional perdurance of Cas9 activity might create an unwanted color progression in progenies (Fig. 2c). Therefore, at least 50 fly brains are required to achieve accurate reconstruction. Nevertheless, it is still much more efficient than other imaging-based methods. The authors also proposed to enhance SSA effectiveness by incorporating more repeats for SSA and to speed up cascade progressions by a more robust control of Cas9 expression.
Why I liked the preprint :
CLADES is the first imagining based method to trace temporal events of a cell lineage, including changes in morphology and distribution. This elegant design takes the lineage tracing research another step forward. I am particularly impressed by their persistence and rigorous tests to make the CLADES progression works! Besides, CLADES can be customized to study various biological events of interest, as the capacity (number of colors) and resolution (the speed of color progression) can be programmed. Given that the optimized gRNA structure works in zebrafish as well, I look forward to seeing the application of CLADES in vertebrate model systems.
Questions for the authors:
Interestingly, the progression of the CLADES cascade is much faster in Dpn-GAL4 (fig 4B) than GH146-GAL4 (Fig4D) lines (the same for comparing GH146-GAL4 to Acj-GAL4). Does driver activity of the reporter also affect the expression level of embedded gRNAs, therefore influencing the progression rate of the cascade? Is it possible to use a stronger promoter for gRNA expression to speed up reporter progression?
References:
- Ma, J., et al., Neural lineage tracing in the mammalian brain. Curr Opin Neurobiol, 2018. 50: p. 7-16.
- J. Garcia-Marques et al., Unlimited genetic switches for cell-type specific manipulation. bioRxiv (2018), available at (https://www.biorxiv.org/content/10.1101/470443v1)
- Almeida, R. CaSSA: a new versatile system for intersectional gene expression enabling cell-type specific manipulations in vivo. Nov 14, 2018, available at (https://prelights.biologists.com/highlights/unlimited-genetic-switches-for-cell-type-specific-manipulation/ )
Posted on: 7 July 2019 , updated on: 9 July 2019
doi: https://doi.org/10.1242/prelights.11764
Read preprintThank you for your explanation. I’ve incorporated your response into the prelight post. I look forward to advanced versions of CLADES for further applications.
Have your say
Sign up to customise the site to your preferences and to receive alerts
Register hereAlso in the developmental biology category:
Alteration of long and short-term hematopoietic stem cell ratio causes myeloid-biased hematopoiesis
Temporal constraints on enhancer usage shape the regulation of limb gene transcription
OGT prevents DNA demethylation and suppresses the expression of transposable elements in heterochromatin by restraining TET activity genome-wide
Also in the genetics category:
Temporal constraints on enhancer usage shape the regulation of limb gene transcription
A long non-coding RNA at the cortex locus controls adaptive colouration in butterflies
AND
The ivory lncRNA regulates seasonal color patterns in buckeye butterflies
AND
A micro-RNA drives a 100-million-year adaptive evolution of melanic patterns in butterflies and moths
A revised single-cell transcriptomic atlas of Xenopus embryo reveals new differentiation dynamics
Also in the molecular biology category:
Fetal brain response to maternal inflammation requires microglia
Clusters of lineage-specific genes are anchored by ZNF274 in repressive perinucleolar compartments
Nanos2+ cells give rise to germline and somatic lineages in the sea anemone Nematostella vectensis
preLists in the developmental biology category:
BSDB/GenSoc Spring Meeting 2024
A list of preprints highlighted at the British Society for Developmental Biology and Genetics Society joint Spring meeting 2024 at Warwick, UK.
| List by | Joyce Yu, Katherine Brown |
GfE/ DSDB meeting 2024
This preList highlights the preprints discussed at the 2024 joint German and Dutch developmental biology societies meeting that took place in March 2024 in Osnabrück, Germany.
| List by | Joyce Yu |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
2nd Conference of the Visegrád Group Society for Developmental Biology
Preprints from the 2nd Conference of the Visegrád Group Society for Developmental Biology (2-5 September, 2021, Szeged, Hungary)
| List by | Nándor Lipták |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
Single Cell Biology 2020
A list of preprints mentioned at the Wellcome Genome Campus Single Cell Biology 2020 meeting.
| List by | Alex Eve |
Society for Developmental Biology 79th Annual Meeting
Preprints at SDB 2020
| List by | Irepan Salvador-Martinez, Martin Estermann |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
Cell Polarity
Recent research from the field of cell polarity is summarized in this list of preprints. It comprises of studies focusing on various forms of cell polarity ranging from epithelial polarity, planar cell polarity to front-to-rear polarity.
| List by | Yamini Ravichandran |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EDBC Alicante 2019
Preprints presented at the European Developmental Biology Congress (EDBC) in Alicante, October 23-26 2019.
| List by | Sergio Menchero et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |
BSCB/BSDB Annual Meeting 2019
Preprints presented at the BSCB/BSDB Annual Meeting 2019
| List by | Dey Lab |
Zebrafish immunology
A compilation of cutting-edge research that uses the zebrafish as a model system to elucidate novel immunological mechanisms in health and disease.
| List by | Shikha Nayar |
Also in the genetics category:
BSDB/GenSoc Spring Meeting 2024
A list of preprints highlighted at the British Society for Developmental Biology and Genetics Society joint Spring meeting 2024 at Warwick, UK.
| List by | Joyce Yu, Katherine Brown |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
Semmelweis Symposium 2022: 40th anniversary of international medical education at Semmelweis University
This preList contains preprints discussed during the 'Semmelweis Symposium 2022' (7-9 November), organised around the 40th anniversary of international medical education at Semmelweis University covering a wide range of topics.
| List by | Nándor Lipták |
20th “Genetics Workshops in Hungary”, Szeged (25th, September)
In this annual conference, Hungarian geneticists, biochemists and biotechnologists presented their works. Link: http://group.szbk.u-szeged.hu/minikonf/archive/prg2021.pdf
| List by | Nándor Lipták |
2nd Conference of the Visegrád Group Society for Developmental Biology
Preprints from the 2nd Conference of the Visegrád Group Society for Developmental Biology (2-5 September, 2021, Szeged, Hungary)
| List by | Nándor Lipták |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Also in the molecular biology category:
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |











 (No Ratings Yet)
(No Ratings Yet)
5 years
Jorge Garcia-Marques
Thanks Ying-Tsen Tung for selecting our work for your highlight. While we think additional experiments will be required to truly prove that different drivers influence the speed of the cascade, this is a very reasonable argument. Depending on the driver, transcription could occur at different levels or not occur at all. This transcription could interfere with the U6 promoters and therefore reduce the cascade speed. Similar mechanisms of transcriptional interference have been previously described. On the other hand, some U6 genes are naturally located within introns, similar to what we designed in our construct. Moreover, U6 promoters are quite strong and we do not think this is the limiting factor in the cascade. Increasing the Cas9 concentration specifically in neuroblasts or using more accessible loci to integrate the CLADES constructs will probably result in a faster cascade.