Molecular determinants of the Ska-Ndc80 interaction and their influence on microtubule tracking and force-coupling
Posted on: 24 June 2019 , updated on: 17 September 2019
Article now published in eLife at http://dx.doi.org/10.7554/eLife.49539
Ndc80 and Ska: Shedding light on the molecular mechanisms of kinetochore-microtubule attachment and force-coupling in microtubule dynamicity
Selected by Yasmin LauBackground
The correct segregation of chromosomes from mother to daughter cells relies on dynamic chromosome-microtubule attachment. This event during cell division (mitosis or meiosis) enables separation of chromatids towards spindle poles and subsequently, the microtubule lattice. Kinetochores are structures which are comprised of a complex of proteins and are localized on the centromere. These act as binding sites for microtubule-chromosome attachment (1). Chromosome-microtubule binding primarily involves 2 steps: First, the kinetochore attaches to the lattice of the microtubule laterally causing the chromosome to lie horizontally. Second, the chromosome rotates and stands perpendicular to the microtubules, which is followed by end-on attachment of the plus ends of the microtubules to the kinetochores (2). This ultimately allows chromosomes to be pulled in opposite directions and get recruited to the respective cells via kinetochore-plus-end-attachment. This process is called “end-conversion” and is essential for cell survival as well as correct chromosomal configuration. However, so far, its mechanism is poorly understood.
Recent advances have revealed 2 complexes, Ndc80 and Ska, that could play major roles in the end-conversion process (3). Within the Ndc80 structure (Figures a and b), microtubule binding regions in Ndc80 include a globular structure formed by 2 calponin-homology domains towards the N-terminal, as well as a basic tail before the CH domain. Ndc80 is also docked onto the kinetochore via a C-terminal (SPC25 and SPC24) RWD domain (4)(5)(6).
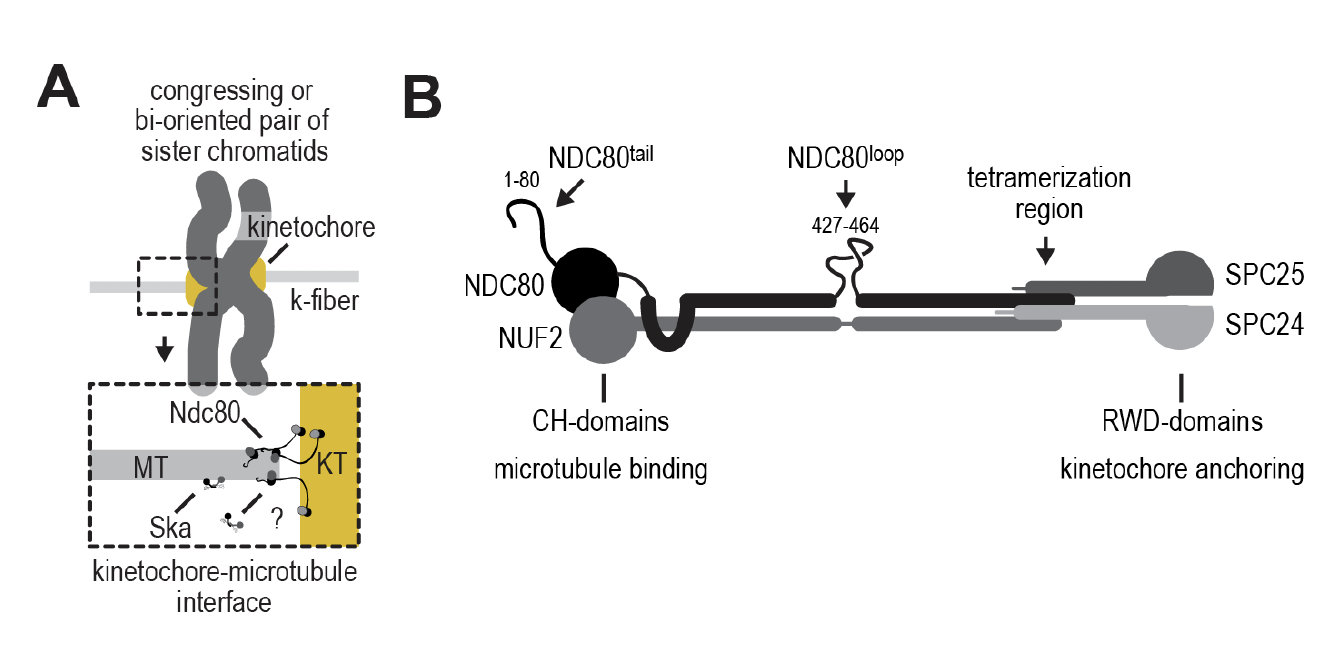
Figure 1a) and b): Ndc80 forms the interface of kinetochore microtubule attachment. The Ndc80 structure involves 2 CH-domains and 2 RWD-domains.
The Ska complex has also been demonstrated to stabilize kinetochore-microtubule attachment. Similar to Ndc80 depletion, absence of Ska has also been shown to cause alignment defects, as well as weak kinetochore fibres (7). However, interestingly, the spindle assembly checkpoint (a cell cycle checkpoint) is activated upon Ska depletion while an opposite phenotype is observed for Ndc80 depletion (8). This difference could shed light on the mechanism of action of Ndc80 and Ska on kinetochores in relation to spindle assembly. Moreover, previous studies have shown that recruitment of Ska to the kinetochore requires Ndc80 (9).
In this preprint, the authors dissect the molecular mechanisms of Ndc80 and Ska in relation to microtubule end-on attachment with kinetochores and force-coupling.
Key Findings
Phosphorylation of SKA3 via CDK1:Cyclin B mediates Ska binding to Ndc80
As the Ska complex has been implicated to be phosphorylated in mitosis by Aurora B, MPS1 and CDK1, the effect of this phosphorylation on Ndc80 binding was sought after. Among the 3 kinases, suppression of CDK1 was found to dampen Ska phosphorylation; CDK1-dependent phosphorylation in vitro was also verified with mass spectrometry, in which 12 of 14 CDK sites were phosphorylated. Moreover, binding to Ndc80 upon Ska phosphorylation was tested through interactions between purified Ndc80 and Ska complexes with SEC. It was found that phosphorylated Ska eluted earlier when mixed with Ndc80 demonstrating that a larger complex is formed. Furthermore, treatment of post-phosphorylated Ska with l-phosphatase prevented complex formation and caused later elution. Taken together, these results indicate that Ska binds Ndc80 upon phosphorylation of the SKA3 subunit.
Ska:Ndc80 binding in vitro is not affected by Aurora B kinase
As previous studies have shown that Aurora B kinase prevents recruitment of Ska to kinetochores, and Aurora B kinase has been shown to phosphorylate the SKA1 and SKA3 subunits, the authors asked whether it also regulates Ska interaction with Ndc80. Firstly, stability of the Ska:Ndc80 complex was not affected upon exposure to Aurora B kinase; this was confirmed with elution times of both Ndc80 and Ndc80:Ska in exposure to Aurora B, as well as with the migration of this complex in SDS-PAGE. These results demonstrate that while Aurora B kinase can prevent recruitment of Ska to the kinetochore, it does not affect Ska:Ndc80 binding.
Ska stabilization of end-on Ndc80-microtubule interactions is force-driven
Next, the authors asked whether force-coupling is influenced by Ska binding. This was approached by exposing both phosphorylated and unphosphorylated Ska to glass beads coated with trivalent Ndc80 in force experiments. The purpose of this is to assess association of both forms of Ska against the dynamicity of the microtubule lattice influenced by force-induction (Figure 2).
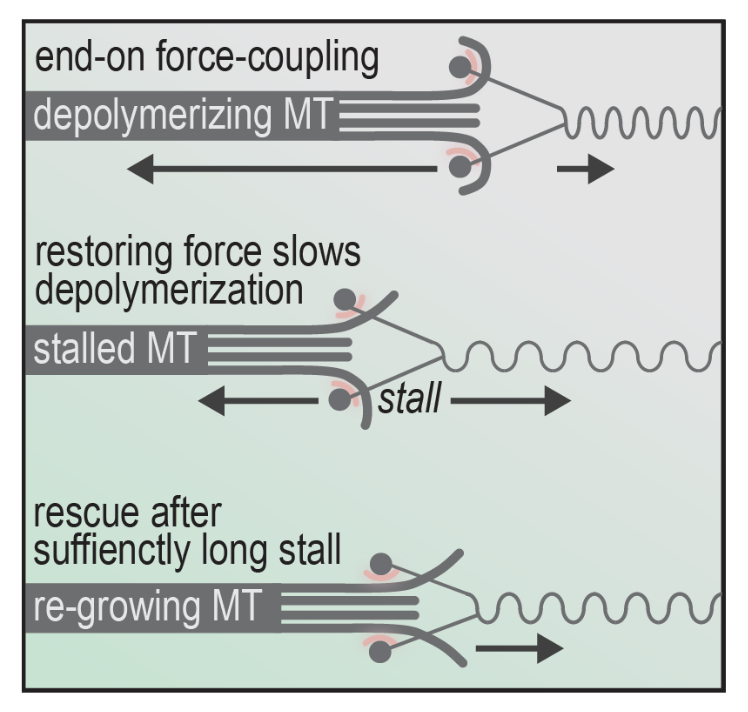
Figure 2: The mechanism of changing microtubule dynamics between depolymerization, stalling and re-growing leading to bi-orientation.
Specifically, stalling force was measured against duration of stalls with and without Ska in conjunction with different Ndc80-tail constructs (such as unphosphorylated and phosphorylated). The authors found that the presence of Ska significantly increased stall duration at the same force in pN, showing the ability of Ska to stabilize microtubules during force-induced stalling. This is interesting, as additional players independent of the kinetochore such as Ska can affect the dynamicity of microtubule growing and shrinking.
Why I like this preprint
Chromosome segregation in mitosis is heavily influenced by force-coupling mechanisms of microtubules, as well as dynamics of kinetochore-microtubule attachment. Cell cycle checkpoints are also regulated by environmental cues associated to these dynamics. Errors in such mechanisms can therefore result in detrimental effects on both chromosomal inheritance and cell survival. It is also intriguing to investigate the dynamics of kinetochore-microtubule attachment from a biophysical point of view, as this preprint partially questions the effect of force inflicted on microtubules by additional cargo which influences their motility, and the limit of this force that microtubules can undergo during chromosome attachment.
Questions
In light of the force experiment using glass beads, how reflective is this experimental design of the actual kinetochore-microtubule mechanism? Could there be other players within the kinetochore that also influence microtubule dynamicity that are not present in the experiment?
References
- Musacchio A., and Desai A., (2017). A molecular view of kinetochore assembly and function. Biology (Basel), 6.
- Dong Y., Vanden Beldt. K.J., Meng X., Khodjakov A., and McEwan B.F., (2007). The Outer Plate in Vertebrate Kinetochores is a flexibile network with multiple microtubule interactinos. Nat Cell Biol, 9, 516-522.
- Auckland P., and McAinsh A.D., (2015). Congressing kinetochores progressively load Ska complexes to prevent force-dependent detachment. J Cell Biol, 216, 1623-1639.
- Alushin G.M., Musinipally V., Matson D., Tooley J., Stukenberg P.T., and Nogales E., (2012). Multimodal microtubule binding by the Ndc80 kinetochore complex. Nat Struct Mol Biol, 19, 1161-1167.
- Cheerambathur D.K., Prevo B., Hattersley N., Lewellyn L., Corbett K.D., Oegema K., and Desai A., (2017). Dephosphorylation of the Ndc80 tail stabilizes kinetochore-microtubule attachments via the Ska complex. Dev Cell, 41, 424-437 e424.
- Cheeseman I.M., Anderson S., Jwa M., Green E.M., Kang J., Yates J.R. 3rd, Chan C.S., Drublin D.G., and Barnes G., (2002). Phospho-regulation of kinetochore-microtubule attachments by the Aurora kinase Ipl1p. Cell, 111, 163-172.
- Daum J.R., Wren J.D., Daniel J.J., Sivakumar S., McAvoy J.N., Potapova T.A., and Gorbsky G.J., (2009). Ska3 is required for spindle checkpoint silencing and the maintenance of chromosome cohesion in mitosis. Curr Biol, 19, 1467-1472.
- Kim S., and Yu H., (2015). Multiple assembly mechanisms anchor the KMN spindle checkpoint platform at human mitotic kinetochores. J Cell Biol, 208, 181-196.
- Chan Y.W., Jeyaprakash A.A., Nigg E.A., and Santamaria A., (2012). Aurora B controls kinetochore-microtubule attachments by inhibiting Ska complex-KMN network interaction. J Cell Biol, 196, 563-571.
doi: https://doi.org/10.1242/prelights.11537
Read preprint










 (No Ratings Yet)
(No Ratings Yet)