Non-neuronal expression of SARS-CoV-2 entry genes in the olfactory system suggests mechanisms underlying COVID-19-associated anosmia
Posted on: 28 April 2020 , updated on: 29 April 2020
Preprint posted on 9 April 2020
Article now published in Science Advances at http://dx.doi.org/10.1126/sciadv.abc5801
How does COVID19 take away your sense of smell? It’s not the neurons in the nose and brain but the support cells that are likely to cause anosmia in COVID-19 patients.
Selected by Sejal DavlaIt started as anecdotal information where doctors and coronavirus patients reported disturbances in the perception of smell and taste leading to anosmia, hyposmia, ageusia, and/or dysgeusia. Now, the loss of sense and/or taste as a symptom of COVID-19 infection is recognized by the medical community worldwide and any adult with a loss of olfactory sense is now asked to self-isolate despite not showing any other symptoms. This post-COVID anosmia is now responsible for up to 40% of cases of anosmia1.
A recent preprint from Brann et al explores the putative mechanism behind how CoV-2 might alter smell and taste perception in patients using RNA sequencing datasets from mouse, primate, and human nasal epithelia.
Background
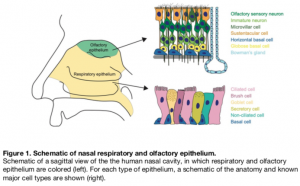
The nasal epithelium constitutes the uppermost part of the respiratory system and harbors respiratory epithelium and sensory olfactory epithelium, both of which differ in cell types and function. Humans only have a small proportion of olfactory epithelium whereas in mouse half of the nasal epithelium is made up of olfactory epithelium. The olfactory epithelium is the primary smell sensing tissue that contains olfactory sensory neurons (OSNs) expressing receptors to detect odorant molecules. The OSN terminals end in the olfactory bulb in the brain where the smell signal is further processed.
The OSNs receive structural support by sustentacular cells in the olfactory epithelium that also maintain salt/water balance and clear toxic agents. The olfactory epithelium also harbors a specialized basal stem cell population that renews OSNs and sustentacular cells throughout life. Among the stem cells, the globose basal cells renew OSNs whereas horizontal basal cells are activated upon tissue damage to regenerate epithelium.
Viruses use a set of receptors and proteases to identify and infect host cells via membrane fusion. SARS-CoV-2 employs the viral spike (S) protein to detect receptor protein ACE2 on target cells. The cleavage of S protein utilizes another host protein – cell surface protease TMPRSS2 to activate the membrane fusion function of S protein. Sequencing datasets show high levels of ACE2 and TMPRSS2 in the nasal respiratory epithelium suggesting respiratory epithelium cells may act as a virus reservoir during CoV2 infections2.
Do cells in olfactory epithelium express proteins for CoV2 entry which makes them prone to direct infections? In light of how the virus infects human cells, whether anosmia is the primary result of olfactory epithelium cells under attack is investigated in this manuscript.
Key findings
The RNA sequencing (RNAseq) datasets identified molecules responsible for CoV2 infections enriched in mouse nasal epithelium. The smell sensing OSNs in both bulk and single-cell RNAseq (scRNAseq) datasets did not show enrichment of Ace2 transcripts making them less likely to be a direct target of CoV2 infection. Ace2 and Tmprss2 expression were largely detected in sustentacular cells, globose basal cells, and horizontal basal cells. The olfactory bulb neurons that synapse with OSNs in the brain do not express Ace2. However, the vascular pericytes in the olfactory bulb that form the blood-brain barrier and trigger inflammatory responses are Ace2 positive.
The authors further probed all CoV2 entry related genes in the sequencing data from the whole olfactory mucosa of primates and humans. Similar to mouse datasets, scRNAseq from nasal biopsy samples in humans did not detect ACE2 and TMPRSS2 in mature OSNs but only in sustentacular and horizontal basal cells. ACE2 immunohistochemistry on human olfactory mucosal biopsy samples revealed localization on sustentacular and basal cells but not on OSNs. Contrary to CoV2, genes for the entry of other CoVs are enriched in both OSNs and support cells.
These results highlight a putative mechanism of olfactory dysfunction where SARS-CoV2 does not directly infect the olfactory sensory neurons or the neurons in the olfactory bulb, the primary olfactory target in the brain. Instead, the viral assault on neurons’ support cells and stem cells may cause architectural damage to the olfactory system. Similar findings were also reported by Fodoulin et al using the newly generated scRNAseq dataset3.
Furthermore, Brann et al compared the frequency of ACE2 and TMPRSS2 expression in human olfactory epithelium vs respiratory epithelium. All respiratory epithelium cell types had a higher frequency of ACE2 expression compared to cells in the olfactory epithelium. The frequency of ACE2 expression was lower in olfactory horizontal basal cells compared to respiratory counterparts. Some olfactory epithelial cell types show high expression of TMPRSS2 but low frequency of ACE2 such as Microvillar and Bowman’s gland cells. Together, this analysis suggests CoV2 may target respiratory epithelium with a higher frequency than the olfactory epithelium.
What I liked about this study
This study is timely and impactful in addressing the putative mechanism behind one of the most talked-about symptoms of COVID19 – anosmia. The authors used a powerful approach using previously published transcriptomics data to sort out cells in the nasal epithelium that are potential targets of SARS-CoV-2. The authors further corroborated the transcriptome data using immunohistochemistry against Ace2 in the nasal epithelium despite lab closures.
To quote the senior author of this study Dr. Datta – “Brad Goldstein, our collaborator at Duke did the immunohistochemistry. I think he managed to do this during the shutdown. Given the challenges in getting this sort of human tissue in any case (which was his thing entirely in this paper), it was a real miracle that we could get those data”.
doi: https://doi.org/10.1242/prelights.19681
Read preprintHi Liam,
Great question, Liam. Many people have recovered from COVID-19 associated anosmia but we don’t have numbers on what proportion.
How this recovery relates to ACE2 expression in NE cell types is a pressing question. Please see the Author’s response to question 1. Knowing that COVID19 mostly affects support cells and stem cells, to what extent these cells are damaged in patients and how they regenerate after the viral assault will directly address your question. Unfortunately, no such report is available as of yet.
Also, it remains to be identified whether COVID patients develop permanent anosmia.
PS: I accidentally clicked the Report button on your comment so if you receive any notification, please ignore. Sorry about that.











 (4 votes)
(4 votes)
6 years
Liam Hurst
Does this mean people suffering with COVID-19 associated anosmia will fully recover their smell eventually?