Pal stabilises the bacterial outer membrane during constriction by a mobilisation-and-capture mechanism
Posted on: 22 October 2019
Preprint posted on 4 October 2019
How to pull in the last layer: TolA, TolB and Pal coordinate the outer membrane septation during cell division.
Selected by Hannah BehrensBackground
Gram-negative bacteria have a complex cell envelope consisting of an inner membrane, the periplasm containing the peptidoglycan cell wall, and the outer membrane (OM). The division of all these layers must be coordinated carefully to avoid rupturing the cell. The constriction of the inner membrane and restructuring of the cell wall are coordinated by the divisome which is established by a ring of FtsZ running around the septum. How constriction of the OM is achieved remains poorly understood. A further complication is that there is no source of energy at the outer membrane, as ATP is only present in the cytoplasm and the proton motif force (PMF) runs across the inner membrane. The preprint by Szczepaniak et al. explains an elegant model of how constriction is coordinated with septation.
Involved are three proteins: Pal, which connects OM and cell wall, TolB, a soluble protein in the periplasm that can bind Pal thereby removing it from the cell wall, and TolA, an inner membrane protein that can bind TolB.
Findings and interpretation
- With the onset of cell division there is a 50% increase in the number of Pal molecules at the septum.
- Pal diffuses very slowly in non-dividing cells. In dividing cells, it diffuses more quickly throughout the cell with exception of the septum, where it diffuses at similar speed as in non-dividing cells.
- Required for the faster diffusion are TolA, TolA coupling to the PMF, and TolB binding to Pal.
- The structure of TolA reveals that it works by a similar mechanism as the force-transducing protein TonB, which uses energy from the PMF to remove plug domains from nutrient transporters in the outer membrane.
- Molecular dynamics simulations and mutation data suggest that TolB is force labile, like the plugs that TonB removes from nutrient transporters.
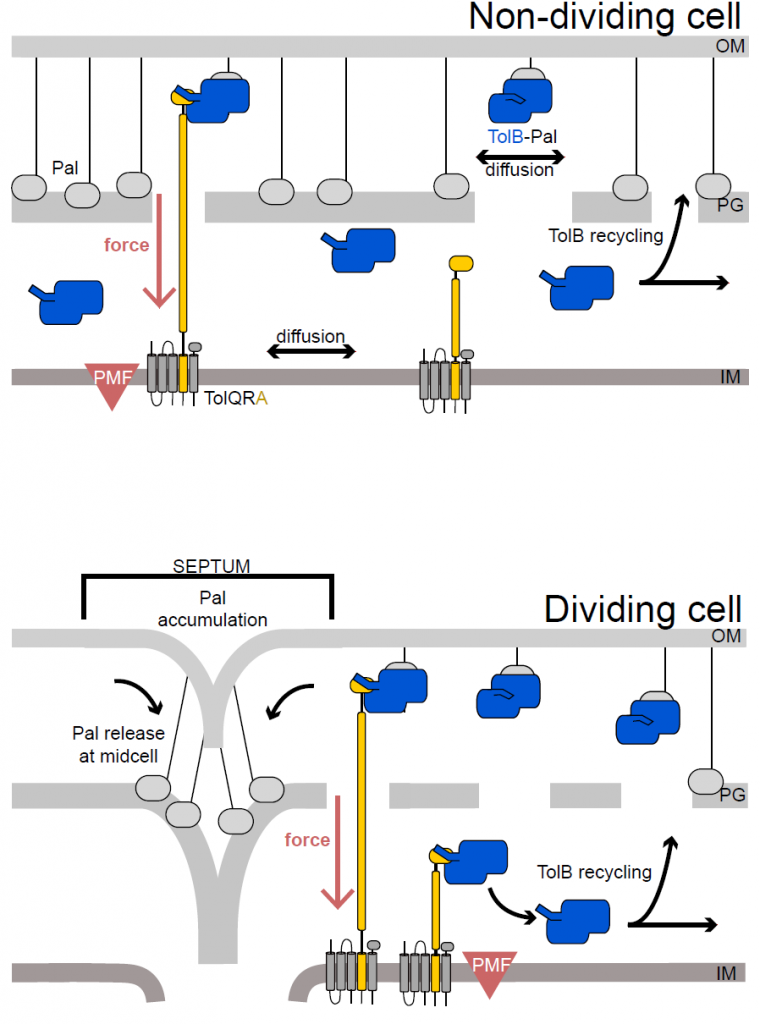
Figure 5 from the preprint: In dividing cells TolA (yellow) is distributed all over the inner membrane, while in dividing cells TolA is located at the septum. As a result, Pal is transported to the septum of dividing cells by TolB and forms a connection between OM and cell wall where TolA is located. This stabilizes the OM during cell division.
The authors combine these finding into a model suggesting that in non-dividing cells most Pal is bound to the peptidoglycan of the cell wall. Occasionally TolB, of which there is only one for every ten molecules of Pal, binds to Pal, thereby displacing it from the peptidoglycan. The TolB-Pal complex is able to diffuse until TolA, which continuously scans the membrane for such complexes, binds to TolB and uses force from the PMF to dissociate the complex. Pal binds to the cell wall again in its new position, while TolB, now on the inner side of the cell wall, diffuses around the periplasm until it finds a new Pal molecule. This mechanism means that Pal is most likely to form stabilizing links between outer membrane and peptidoglycan where TolA is located.
In dividing cells TolA is located to the septum. As a result, TolB displaces Pal form peptidoglycan throughout the cell and the TolB-Pal complex diffuses throughout the cell like before. But in dividing cells, TolA is only at the septum where it removes TolB from Pal, allowing Pal to bind to peptidoglycan. By this process, which the authors term mobilization-and-capture mechanism, Pal is accumulated at the septum where it forms a stable connection between the outer membrane and newly formed peptidoglycan. This stabilization is needed to prevent membrane damage during cell division.
What I like about this preprint and future directions
There are two questions in the field of the Gram-negative cell envelope that are addressed by the model proposed by Szczepaniak et al. The first question is about the mechanism of the tol-phenotype, discovered in the 60s. Deletion of tol genes, like tolA and tolB, lead to OM instability, hypersensitivity towards detergents and bile salts, and leakage of periplasmic contents. For over forty years it has been unknown how the deletion of tol genes leads to these defects.
The second question is a result of recent progress in the understanding of the mechanisms that coordinate bacterial division. FtsZ monomers form a treadmilling ring around the division site, onto which the divisome, a large complex involved in septation, is assembled. Recent papers have outlined how inner membrane septation and cell wall remodelling are regulated but how these events are coordinated with the constriction of the OM remained largely unknown.
The mobilization-and-capture mechanism reveals that both questions have the same answer, that the components of the Tol-system are essential in coordinating OM constriction, thereby preventing membrane damage which would cause a tol-phenotype.
I am curious to see whether future research will uncover what drives TolA localisation to the septum.
References/Further reading
Bisson Filho, Alexandre Wilson, Yen-Pang Hsu, Georgia Squyres, Erkin Kuru, Fabai Wu, Calum Jukes, Cees Dekker, et al. 2017. “Treadmilling by FtsZ Filaments Drives Peptidoglycan Synthesis and Bacterial Cell Division.” Science 355 (February): 739–43. https://doi.org/10.1101/077560.
Petiti, Mélissa, Bastien Serrano, Laura Faure, Roland Lloubes, Tâm Mignot, and Denis Duché. 2019. “Tol Energy-Driven Localization of Pal and Anchoring to the Peptidoglycan Promote Outer-Membrane Constriction.” Journal of Molecular Biology 431 (17): 3275–88. https://doi.org/10.1016/j.jmb.2019.05.039.
Yang, Xinxing, Zhixin Lyu, Amanda Miguel, Ryan McQuillen, Kerwyn Kerwyn Casey Huang, and Jie Xiao. 2017. “GTPase Activity-Coupled Treadmilling of the Bacterial Tubulin FtsZ Organizes Septal Cell-Wall Synthesis.” Science 355 (February): 744–47. https://doi.org/10.1101/077610.
doi: https://doi.org/10.1242/prelights.14768
Read preprint










 (No Ratings Yet)
(No Ratings Yet)