Live imaging of Aiptasia larvae, a model system for studying coral bleaching, using a simple microfluidic device
Posted on: 9 August 2018 , updated on: 12 August 2018
Preprint posted on 19 July 2018
Article now published in Scientific Reports at http://dx.doi.org/10.1038/s41598-019-45167-2
How can one image motile sea anemone larvae? A recent preprint by Van Treuren and Brower et al. introduces ‘Traptasia’, a microfluidic device capable of trapping and imaging live Aiptasia larvae, a model for coral symbiosis.
Selected by Samantha SeahCategories: ecology
Background
Coral reefs are vital ecosystems in our oceans that are home to a tremendous diversity of oceanic species. Due to environmental stressors, along with stress caused by humans, corals are increasingly undergoing ‘bleaching’ – a process in which photosynthetic algal symbionts are ejected from the corals. Over the long term, this could lead to coral death. Even as the stress factors contributing to coral bleaching are known, much work remains to be done to thoroughly elucidate the molecular and cellular mechanisms involved.
Coral symbiosis can be modelled by studying the motile larvae of sea anemone, Aiptasia, together with symbiotic algae from the genus Symbiodinium. A recent preprint by Van Treuren, Brower and their colleagues outline the trapping and imaging of motile Aiptasia larvae, via a microfluidic device, ‘Traptasia’, enabling dynamic observations under different conditions.
Key Findings
The authors designed a single layer polydimethylsiloxane (PDMS) trapping device capable of capturing Aiptasia larvae. These resemble previously described cell traps, but as Aiptasia larvae (at 40-100µm) are much larger than typical mammalian cells (around 10µm), the authors optimised and assessed trap-loading efficiencies for traps with varying trap apertures and channel heights. The optimised traps (with heights of 90µm and trap apertures of 20µm) were able to efficiently capture Aiptasia larvae and hold the larvae in traps by the provision of a constant flow of fluid.
The authors then demonstrated that trapped larvae can be imaged by spinning disk confocal microscopy for over 10 hours. Larvae co-infected with algal symbionts could also be imaged with transmitted light alone, and individual algae tracked within the larvae. Larval revolutions could also be followed, enabling the further study of subtle stress responses. The authors also noted multiple potentially interesting larval-death mechanisms which had been previously observed in culture, but have yet to be studied in cellular detail, illustrating how the ‘Traptasia’ could reveal mechanistic details of Aiptasia physiology.
To model coral bleaching upon stress, the authors treated algae-larvae symbionts with DCMU (3-(3,4-dichlorophyli)-1,1-dimethylurea), which has been proposed to stress corals. Under this treatment, 5/33 trapped larvae ‘swam’ upstream of flow or through the trap aperture, suggesting that stress may introduce changes in motility and physiology. In addition, they captured an algal expulsion event from a live Aiptasia larva (Video S6). As algal expulsion is thought to be involved in coral bleaching, the use of ‘Traptasia’ to study Aiptasia larvae under different stress conditions could reveal biological mechanisms involved in coral bleaching, potentially shedding light on the damage accumulating in our coral reefs.
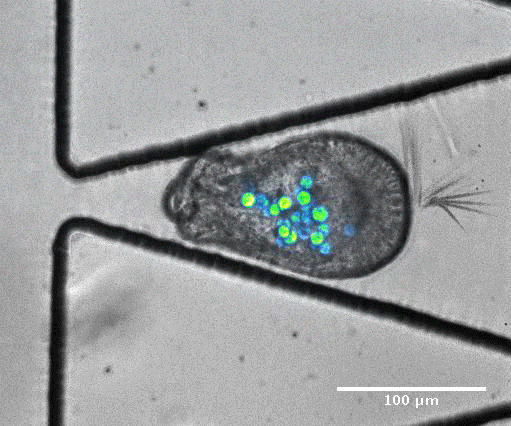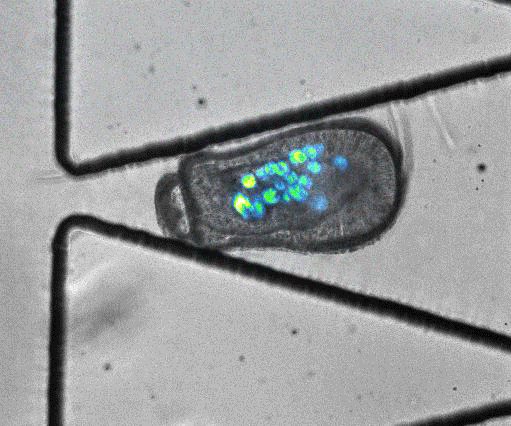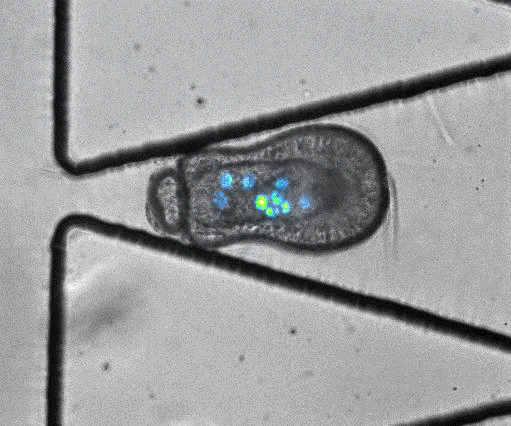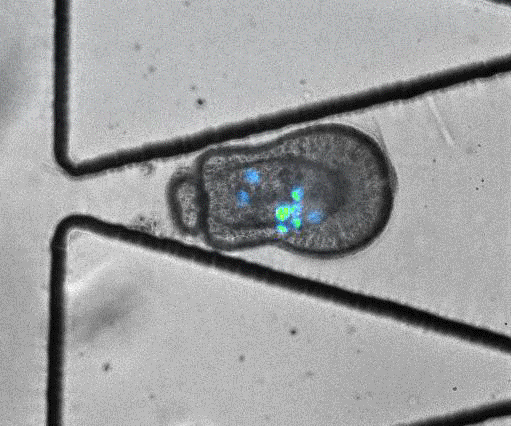
Video S6 from the preprint: Algal expulsion event from live Aiptasia larva. Video reproduced under a CC-BY-NC-ND 4.0 International License.
What I like about this work
As a biologist working in the field of microfluidics, I am a great fan of simple microfluidic devices that can be used by biologists to complement the study of otherwise complex biological problems. ‘Traptasia’ falls neatly into this category – a single layer microfluidic device is used to trap and image large, deformable and motile organisms, without the need for excessively complex equipment. As individual algae can be imaged with transmitted light, this technology can be made accessible to many locations, including schools and field research stations.
I see so much potential in this new technology – combining it with genomics and transcriptomics (as mentioned by the authors) could add dimensionality to the data obtained. It could be also adapted for use with other marine creatures and motile organisms, and I am keen to see how others will adapt this technology to investigate other biological problems.
Open Questions
- Do you see the traps working for heterogenous populations, and if not, would it be possible to design such traps?
- Could the devices be utilised for the short-term culture and study of primary aquatic samples?
Further reading:
Microfluidic cell trapping: Dura, B., Servos, M. M., Barry, R. M., Ploegh, H. L., Dougan, S. K., & Voldman, J. (2016). Longitudinal multiparameter assay of lymphocyte interactions from onset by microfluidic cell pairing and culture. Proceedings of the National Academy of Sciences, 113(26), E3599–E3608. https://doi.org/10.1073/pnas.1515364113
Coral-on-a-chip: Shapiro, O. H., Kramarsky-Winter, E., Gavish, A. R., Stocker, R., & Vardi, A. (2016). A coral-on-a-chip microfluidic platform enabling live-imaging microscopy of reef-building corals. Nature Communications, 7, 10860. https://doi.org/10.1038/ncomms10860
doi: https://doi.org/10.1242/prelights.4182
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the ecology category:
Feeding and reproduction of a tropical coastal copepod across warming and copper gradients
Tina Nguyen
Heatwave winners and losers: cryptic coral holobionts differ in thermal tolerance
Maitri Manjunath
No global collapse of food webs across the Permian–Triassic Mass Extinction
Theodora Stougiannou
preLists in the ecology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
EMBO | EMBL Symposium: The organism and its environment
This preList contains preprints discussed during the 'EMBO | EMBL Symposium: The organism and its environment', organised at EMBL Heidelberg, Germany (May 2023).
| List by | Girish Kale |
Bats
A list of preprints dealing with the ecology, evolution and behavior of bats
| List by | Baheerathan Murugavel |











 (1 votes)
(1 votes)