Agrobacterium tumefaciens divisome proteins regulate the transition from polar growth to cell division
Posted on: 20 September 2018
Preprint posted on 10 September 2018
Article now published in Molecular Microbiology at http://dx.doi.org/10.1111/mmi.14212
No matter whether polar or lateral growth, FtsZ rings drive the assembly of the divisome to coordinate bacterial cell division.
Selected by Hannah BehrensCategories: microbiology
Background
While the mechanisms of eukaryotic cell division have been worked out in great detail (and has made it to the textbooks), we are just starting to understand the molecular mechanisms of bacterial cell division.
Last year, a long standing question was answered: what coordinates cell wall and membrane constriction and ultimately cell division? It was found that a protein called FtsZ, a tubulin-like GTPase, forms constricting rings around the division site. FtsZ then recruits other cell division proteins, and the resulting complex is called the divisome. One of the divisome proteins is peptidoglycan synthase which produces new peptidoglycan to form the cell wall at the division site. This process has been investigated in the model organisms Bacillus subtilis and Escherichia coli, respectively.
Here Howell et al. investigate the division process in Agrobacterium tumefaciens, a bacterium that is of interest for two reasons. First, it belongs to the group of alpha-proteobacteria, Gram negative bacteria in which the role of FtsZ and the other divisome components is largely unstudied. Secondly, it has polar growth, which means the bacteria only grow in one direction: one pole grows, the other one doesn’t. This is opposed to B. subtilis and E. coli which have lateral growth. The different mode of growth posed the question whether cell division is coordinated in the same way in A. tumefaciens as in the previously studied B. subtilis and E. coli.
A. tumefaciens is also an important molecular biology tool in the genetic modification of plants.
Key findings
This paper is a first of its kind characterization of the A. tumefaciens divisome. The effects of depletion of the divisome components FtsZ, PopZ, FtsA, FtsW and the effect of DNA replication are investigated.
- There are three FtsZ homologues in tumefaciens; FtsZAT is the major one, resulting in a striking depletion phenotype shown below, while FtsZ1 appears to be a non-essential divisome component. FtsZ2, the third homologue, does not localize to the division site and was not investigated further.
- FtsZAT forms filaments. FtsZ1 can integrate in FtsZAT filaments, but not form filaments of its own.
- If PopZ or FtsZAT is depleted the respective other component does not localize correctly, indicating both are important for divisome positioning and functioning.
- DNA replication can start but not go to completion without cell division.
- The loss of peptidoglycan synthesis at the division site in FtsZ depletion results in altered peptidoglycan composition.
- The CTP domain of FtsZAT coordinates termination of polar growth, the CTL domain is required for correct peptidoglycan composition and protofilament assembly.
- FtsA is required for cell division, deletion of the downstream component FtsW has the same effect as deletion of FtsA.
In conclusion, the previous model that FtsZ rings drive the formation of the divisome to coordinate cell division holds true. How the behaviour of individual divisome components compares to other that in other species is discussed in detail in the preprint and is beyond the scope of this highlight.
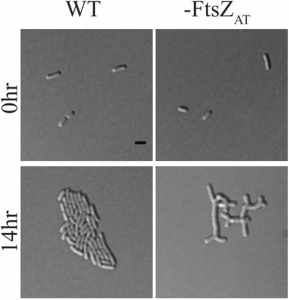
Figure 1D adapted from the preprint: FtsZAT depletion results in branched cells with a large surface area. This is opposed to E. coli and B. subtilis which form long smooth cells when FtsZ is depleted. You can find a movie of it here https://twitter.com/Michelle12Will/status/1039262678353620998.
What I like about this preprint
In a structured manner, the authors investigate and describe the effects that each of the four key components has on the divisome and then compare these to observations made in other species. This work adds several pieces to the puzzle that explains bacterial cell division across species and even kingdoms, as the comparison to chloroplasts and mitochondria is not missed out either.
I think the finding that FtsZ coordination of cell division is conserved across different modes of growth (polar growth and lateral growth) is very striking. This extent of mechanistic conservation also suggests to me that the divisome might be interesting as a drug target.
Future directions: application and further research
As the authors outline in their paper, many questions remain to be answered about the mechanisms of cell division in A. tumefaciens.
- How does the FtsZAT ring lead to termination of polar growth?
- Which proteins contribute to polar peptidoglycan synthesis and what triggers the termination of polar growth?
- How is FtsZAT targeted to the division site?
- And once it is at the division site, how is the FtsZAT -ring stabilized?
Personally, I am most excited to see how FtsZAT might be involved in termination of polar growth.
Future experiments in more different species will continue to elucidate cell division across the spectrum of bacteria and plastids.
References/Further reading
These two papers describe for the first time that it is FtsZ treadmilling that organizes cell wall synthesis, both in Gram positive (Bisson et al.) and in Gram negative (Yang et al) bacteria.
- Bisson Filho, A. W. et al. (2016) ‘Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division.’, Science, 355(February), pp. 739–743. doi: 10.1101/077560.
- Yang, X. et al. (2017) ‘GTPase activity-coupled treadmilling of the bacterial tubulin FtsZ organizes septal cell-wall synthesis’, Science, 355(February), pp. 744–747. doi: 10.1101/077610.
This video show how the authors use live cell microscopy to investigate processes like cell division.
- Howell M.J. et al. (2017) ‘Live cell fluorescence microscopy to observe essential processes during microbial cell growth’, J Vis Exp. https://www.jove.com/video/56497/live-cell-fluorescence-microscopy-to-observe-essential-processes.
This review gives background on polar growing bacteria
- Cameron, T. A., Zupan, J. R. and Zambryski, P. C. (2015) ‘The essential features and modes of bacterial polar growth’, Trends in Microbiology. Elsevier Ltd, 23(6), pp. 347–353. doi: 10.1016/j.tim.2015.01.003.
doi: https://doi.org/10.1242/prelights.4834
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the microbiology category:
Gut microbiome changes over the course of multiple sclerosis differentially influence autoimmune neuroinflammation
Carole Djagang et al.
Human Assembloid Model of Emergent Neurotropic Enteroviruses
Mitchell Sarmie
Cryo-EM reveals multiple mechanisms of ribosome inhibition by doxycycline
Leonie Brüne
preLists in the microbiology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
BioMalPar XVI: Biology and Pathology of the Malaria Parasite
[under construction] Preprints presented at the (fully virtual) EMBL BioMalPar XVI, 17-18 May 2020 #emblmalaria
| List by | Dey Lab, Samantha Seah |
1
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |











 (No Ratings Yet)
(No Ratings Yet)