Rational Design of Minimal Synthetic Promoters for Plants
Posted on: 26 May 2020 , updated on: 1 September 2020
Preprint posted on 14 May 2020
Article now published in Nucleic Acid Research at https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gkaa682/5897334
MynSyns: Construction of synthetic promoters based on transcription factor cis-regulatory elements for predictable expression in plants.
Selected by Facundo RomaniCategories: plant biology, synthetic biology
Context
Promoter sequences are a fundamental component of the molecular biology toolkit. In this work Cai and colleagues put a long-standing issue on the table: most plant biologists use a limited set of promoters to control gene expression since the ’80s. Until now, there have been few studies that dissect widely used promoter sequences in order to understand how they work.
Different kinds of promoters are used to achieve desirable expression patterns or levels, and systems biology heavily relies on a diverse promoter toolkit. For example, small synthetic promoters are easier for cloning and can be useful for the design of complex genetic circuits.
The design of synthetic promoters requires the understanding of the endogenous machinery of transcription factors that modulates gene expression in planta. Advances in the study of transcription factors and their binding motifs could help to create new promoters with desirable characteristics.
Major findings
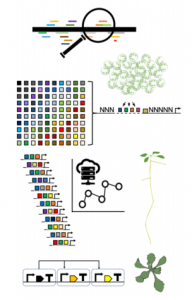
Cai et al. perform a sequence analysis of a series of widely used constitutive promoters. Based on high-throughput analysis of transcription factor binding sites, they use short cis-regulatory DNA elements (CRE) as “parts” for their analysis. As expected, constitutive promoter sequences possess CRE and putative binding sites for many transcription factors. Given that most transcription factors are not expressed constitutively, this implies that promoters are bound by multiple combinations of transcription factors in different contexts. The sequence analysis also identified a regulatory element common to pathogenic promoters (C-CRE).
They developed a transient expression system based on a luminescence reporter that allows quantitative measurement of transcriptional activity for synthetic promoters. In the first place, C-CRE is the main contributor to gene activation in pathogenic promoters and its strength depends on the distance to the transcription start site. They also analyzed other CREs associated with endogenous transcription factors in order to design Minimal Synthetic promoters (MynSyns). Surprisingly, MynSyns with three copies of an identical CRE are not efficient for gene activation while a combination of three different CREs is much more effective. The exception is C-CRE which can deliver high expression levels in multiple copies. They also tested different arrangements of spacers and the positional dependence of CRE contribution to gene activation.
With the knowledge acquired from CRE analysis (eg. position dependency and relative activation strength), they developed a prediction algorithm for MynSyns that yielded a plethora of configurations. They obtained an acceptable prediction of activation strength for MynSyns and developed a series of MynSyns with different constitutive expression levels. Additionally, the authors provide an analysis of how these MynSyns work in stable transgenic expression and in the protoplast from three plant species.
They also created promoters with different strength based on two orthogonal systems: TALE proteins (Brückner et al., 2015) and the synthetic transcription factor Gal4:PhiC31 (Vazquez-Villar et al., 2017). These alternatives provide MynSyns that do not directly depend on the endogenous transcription factors. In this case, expression levels are modulated using different numbers of the same binding site for the orthogonal TF. Finally, this system is used to create synthetic genetic circuits as a proof of concept.
What I like about this preprint
This preprint applies synthetic biology tools to explore many aspects of promoters that are not usually analyzed. Their findings are of broad interest for general plant biology and synthetic biology. The proposed model of “passive cooperativity” between different CRE is really interesting and opens interesting questions for transcription factor biology. Moreover, the large collection of synthetic parts and plasmids in this study will be of use in a great diversity of fields.
Future directions
Constitutive promoters are used very frequently in molecular biology and these new promoters are promising. One of the limitations of pathogenic constitutive promoters that harbor the C-CRE is they are not “completely constitutive”, given that in some tissues, growing conditions or developmental stages the expression is not stable (Holtorf et al., 1995, Somssich, 2019). In MynSyns, high expression level promoters still depend on the C-CRE as in other widely used promoters. Even orthologous systems often rely on C-CRE promoters in at least one of the steps of the genetic circuits. As the authors discuss, C-CRE probably depends on the endogenous bZIP transcription factors that are not expressed constitutively, as most transcription factors. Maybe this is the reason why C-CRE promoters are not ubiquitously expressed. It will be interesting to learn more about the use of other CREs (different from C-CRE) that could provide strong activation with fewer limitations. This is a difficult task that needs a more complex analysis on gene expression of synthetic promoters and requires a deeper understanding of the endogenous transcriptional machinery. In this regard, this preprint provides interesting clues about CREs and how to use them as parts for predictable gene expression.
Questions for the authors
1) One of the aspects that it is not completely clear to me is how the computational design of MynSyns and strength prediction was performed in the case where single CREs does not have a significant effect but in combinations with other CREs does have an effect. Do you compute the effect of the combinations or the contribution of every single CRE?
2) In the preprint, you mention a C-CRE that does not contain a TGA binding site and mention that the positional effect on this case is different. How do you think the TGA affects C-CRE promoters?
3) What are the following steps that you envision for the future of synthetic promoters?
References
Holtorf S, Apel K, Bohlmann H. (1995) Comparison of different constitutive and inducible promoters for the overexpression of transgenes in Arabidopsis thaliana. Plant Mol Biol. 29(4):637-46.
Somssich M. (2019). A short history of the CaMV 35S promoter. PeerJ Preprints 7:e27096v3. https://doi.org/10.7287/peerj.preprints.27096v3
Vazquez-Vilar,M., Quijano-Rubio,A., FernandezDel-Carmen,A., Sarrion-Perdigones,A., OchoaFernandez,R., Ziarsolo,P., Blanca,J., Granell,A. and Orzaez,D. (2017) GB3.0: a platform for plant bio-design that connects functional DNA elements with associated biological data. Nucleic Acids Res. 45:2196–2209.
Brückner,K., Schäfer,P., Weber,E., Grützner,R., Marillonnet,S. and Tissier,A. (2015) A library of synthetic transcription activator-like effectoractivated promoters for coordinated orthogonal gene expression in plants. Plant J. 82:707–16.
doi: https://doi.org/10.1242/prelights.21131
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the plant biology category:
A drought stress-induced MYB transcription factor regulates pavement cell shape in leaves of European aspen (Populus tremula)
Jeny Jose
Actin Counters Geometry to Guide Plant Cell Division
Jeny Jose
The nucleus follows an internal cellular scale during polarized root hair cell development
Jeny Jose
Also in the synthetic biology category:
Enzymatic bromination of native peptides for late-stage structural diversification via Suzuki-Miyaura coupling
Zhang-He Goh
Enhancer cooperativity can compensate for loss of activity over large genomic distances
Milan Antonovic
Discovery and Validation of Context-Dependent Synthetic Mammalian Promoters
Jessica L. Teo
preLists in the plant biology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Also in the synthetic biology category:
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |
Advances in Drug Delivery
Advances in formulation technology or targeted delivery methods that describe or develop the distribution of small molecules or large macromolecules to specific parts of the body.
| List by | Zhang-He Goh |











 (No Ratings Yet)
(No Ratings Yet)