Synthetic mammalian signaling circuits for robust cell population control
Posted on: 3 November 2020
Preprint posted on 3 September 2020
Article now published in Cell at http://dx.doi.org/10.1016/j.cell.2022.01.026
Cells learn a new language! Auxin confers new synthetic communication-channel for cells to communicate with.
Selected by Pavithran RavindranCategories: synthetic biology
Background
One of synthetic biology’s greatest challenges is to engineer cells that can learn from both their environment as well as their neighboring cells to make decisions about what program to execute1. Previous studies have been extremely successful in developing methods to engineer cells to integrate endogenous signaling ligands, such as Sonic Hedgehog2. The development of synthetic Notch receptors that provide completely orthogonal/synthetic communication channels made great progress to the eventual goal, but is limited to cell-to-cell contact-mediated signaling3,4. Very recently, synthetic Notch receptors were used with synthetic ligands (GFP) to program cell decisions, but still required two cells with two separate receptors such that the GFP becomes anchored and displayed on one cell and then activates the adjoining receiver cell5. Therefore, there is still work to be done to develop ligand-based orthogonal communication channels that synthetic biologists can use to engineer cells.
The authors of this preprint developed a method that repurposes the plant hormone auxin to control the expression of user-defined genetic programs. They then utilize this system to perform synthetic quorum sensing, a signaling phenomenon present in bacteria that allows for bacterial cells to learn about local cell density and aids in their decision to divide or not. The authors find that a simple circuit from auxin ligand leading to cell death only transiently limits the size of the population; they instead need a paradoxical circuit where the auxin ligand allows for both cell growth and cell death to ensure escape mutations do not occur in their circuit.
Key Findings
In this preprint, the authors set out to develop a method to use the plant hormone auxin to control the expression of genes in an engineered cell. To do so, they found that they only needed two components: (1) ectopic expression of F-box transport inhibitor response 1 (TIR1) from rice (osTIR1) which recruits E2 ubiquitin ligase for targeted degradation and (2) the auxin inducible degron (AID) attached to genes that the user wants regulated. As a first test, they expressed osTIR1 in mammalian CHO-K1 cells as well as mCherry fused to AID and Blasticidin resistance. With this setup, addition of auxin to the media decreases the expression of mCherry and Basticidin resistance (Figure 1). The latter results in the susceptibility of cells to blasticidin at higher doses of auxin hormone. With this, the authors had set up their receiver cell.
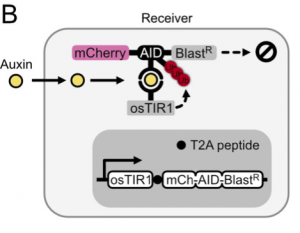
The next step was to develop a way for cells to produce auxin hormone. To do so, they propose that the amino acid tryptophan can be oxidized to a precursor of auxin and that can be hydrolyzed to the final auxin hormone with the help of a hydrolase. After performing a small screen to identify hydrolases that work in mammalian cells, the authors find that by expressing this enzyme, known as iaaH, in their receiver cells, they can simply add precursor to the media and still get degradation of mCherry and Blasticidin (Figure 2). Furthermore, by plating cells containing iaaH (‘sender cells’) on one end of a 60mm dish and filling the rest with receiver cells, the authors find that they get a gradient of mCherry expression in receiver cells where cells near the senders have little to no mCherry fluorescence, whereas those further away have close to unperturbed levels of fluorescence. The authors thus have developed cells that could now send auxin hormone.
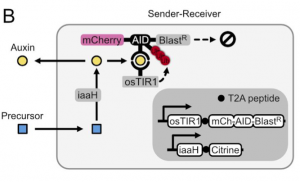
As a proof-of-principle to show the utility of such a system, the authors wanted to allow the cells to sense their own density and regulate their growth based on it. The idea was that by plating sender-receiver cells in the presence of the precursor molecule and Blasticidin, once cells achieved a high enough density, the cells would increase the concentration of auxin in the media resulting in Blast resistance degradation and thus cell death due to susceptibility to Blasticidin. After finding that they needed to express another transport channel (PIN2) to get efficient auxin transport to the media, the authors saw that when they plated the cells at higher density, mCherry fluorescence would decrease. However, when they tried a long time-course of this experiment by plating cells at low density and imaging over time, they found that cells would start initially regulating their confluency in the well, but then start to uncontrollably grow. Upon analysis of the resulting cells that had grown out, the authors found mutations that resulted in auxin-insensitivity and thus constitutive expression of Blasticidin resistance.
Cheater mutations that result in the destruction of genetic circuits are a major problem in synthetic biology. Taking inspiration from previous literature on theoretical circuits that could combat this6, the authors hypothesized that placing an AID on an inducible caspase (iCasp9) and allowing the cells to grow in the small molecule that activates the apoptosis-inducing enzyme will get them around this. With this, the authors have created a paradoxical circuit. At high cell densities with high auxin (a) if the cells have a functioning circuit then Blasticidin resistance is downregulated and cell growth is halted but (b) if cells have broken the circuit (cheater mutation) then iCasp9 is not downregulated leading to the apoptosis of the cheater cell (Figure 3). With this circuit in place the authors show that indeed fewer cheater mutations occur and the cells are better able to control their population size.
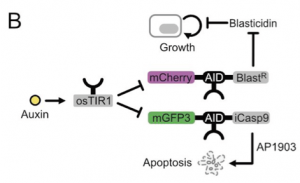
Why I chose this preprint
The development of synthetic channels by which cells can communicate with one another to perform user-defined decisions is a major goal in biology. The use of synNotch juxtacrine signaling has already found many uses in synthetic immunology, but is still limited to cell-to-cell contacts4. Here, the authors of the preprint have found a way to use a small molecule plant hormone (auxin) to drive their genetic circuits. This could open up the possibility for more flexible cellular engineering. Finally, the preprint itself is extremely straightforward to read with each figure containing schematics that clearly delineate the purpose and setup of each experiment.
Questions for the authors
- In most of the current setup, you need to add precursor molecule to drive production of the auxin and turn on the circuit, although you do show in supplementary figure 1 that one can express iaaM to produce the precursor molecule in the sender-receiver cells. Can you comment on why you decided to keep using precursor molecule instead of this setup?
- Is auxin safe in humans? If not, what ideas do you have to port this system for engineered cells for therapeutic cases?
References
- Bashor, C. J. & Collins, J. J. Understanding Biological Regulation Through Synthetic Biology. Annu. Rev. Biophys. 47, (2018).
- Li, P. et al. Morphogen gradient reconstitution reveals Hedgehog pathway design principles. 0645, 1–10 (2018).
- Morsut, L. et al. Engineering Customized Cell Sensing and Response Behaviors Using Synthetic Notch Receptors. Cell 164, 780–791 (2016).
- Roybal, K. T. et al. Engineering T Cells with Customized Therapeutic Response Programs Using Synthetic Notch Receptors. Cell 167, 1–14 (2016).
- Toda, S., Mckeithan, W. L., Hakkinen, T. J. & Lopez, P. Engineering synthetic morphogen systems that can program multicellular patterning. Science (80-. ). 331, 327–331 (2020).
- Hart, Y. et al. Paradoxical signaling by a secreted molecule leads to homeostasis of cell levels. Cell 158, 1022–1032 (2014).
doi: https://doi.org/10.1242/prelights.25551
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the synthetic biology category:
Enzymatic bromination of native peptides for late-stage structural diversification via Suzuki-Miyaura coupling
Zhang-He Goh
Enhancer cooperativity can compensate for loss of activity over large genomic distances
Milan Antonovic
Discovery and Validation of Context-Dependent Synthetic Mammalian Promoters
Jessica L. Teo
preLists in the synthetic biology category:
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |
Advances in Drug Delivery
Advances in formulation technology or targeted delivery methods that describe or develop the distribution of small molecules or large macromolecules to specific parts of the body.
| List by | Zhang-He Goh |











 (No Ratings Yet)
(No Ratings Yet)