Galectin-8 senses phagosomal damage and recruits selective autophagy adapter TAX1BP1 to control Mycobacterium tuberculosis infection in macrophages
Posted on: 25 July 2020 , updated on: 25 March 2021
Preprint posted on 1 July 2020
Article now published in mBio at http://dx.doi.org/10.1128/mBio.01871-20
Guardians from the galectins: Danger sensor galectin-8 promotes degradation of M. tuberculosis using selective autophagy
Selected by Josie Gibson, Zhang-He GohCategories: microbiology
preLight Authors’ note
This is the second preLight of a three-part series on airway inflammation, infection, autophagy and its mediators. Links to the first and third preLights.
Background:
Mycobacterium tuberculosis is an important human pathogen which causes tuberculosis, a disease which results in a large mortality worldwide [1]. M. tuberculosis antibiotic resistance is increasing and present treatments are not ideal. Therefore, there is great interest in understanding host-pathogen interactions in order to identify new therapeutic targets for treatment of tuberculosis.
Autophagy is a cellular self-degradation pathway used to recycle damaged or unwanted cellular components, accomplished through lysosomal degradation. In some cases, autophagy can be used to target specific components, termed selective autophagy, which is completed using autophagy adaptors which bind to both autophagy pathway proteins and ubiquitinated cargo. Autophagy adaptors are present in the cytosol and target ubiquitinated pathogens which have escaped the phagosome and are therefore accessible to cytosolic proteins [2].
M. tuberculosis is phagocytosed by macrophages, it has been reported to damage the vesicle it resides within, in order to escape degradation and enable continued infection. The authors have previously demonstrated that 30% of M. tuberculosis escaped to the cytosol become ubiquitinated [3], suggesting that M. tuberculosis can be targeted by autophagy adaptors. The mechanism of how selective autophagy may be activated in M. tuberculosis infection is not fully understood, so the authors choose to examine a potential role of galectins, which are danger sensors. Cytosolic galectins act to recognise and bind to sites of lysosomal or endosomal membrane damage and are known to interact with other intracellular pathogens as well as autophagy adaptors [4]. In this study, the authors examine how macrophages sense and respond to M. tuberculosis located within damaged phagosomes, with a focus on the mechanistic roles of galectins and selective autophagy (Fig. 1).
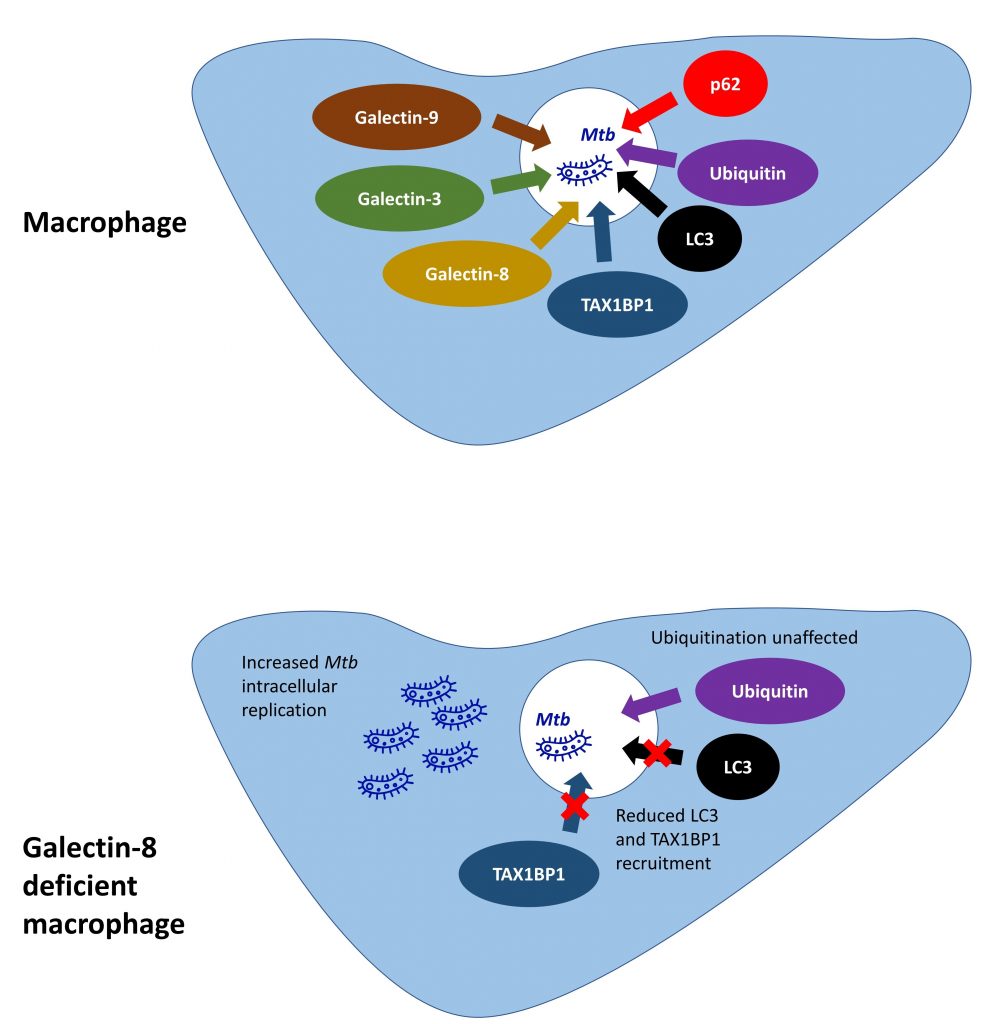
Figure 1. Summary of preprint findings.
Key findings:
The initial important observation in this study is that M. tuberculosis targeted by selective autophagy machinery within macrophages, are also associated with galectin-8. The authors first demonstrate that three different cytosolic galectins, galectin-3, -8 and -9, can be recruited to M. tuberculosis within damaged phagosomes, by expressing fluorescently tagged galectin proteins within macrophages and clearly visualising these co-localising with M. tuberculosis. Next, further imaging shows that M. tuberculosis (in addition to galectin-8) co-localises with ubiquitin, p62 and LC3, all key proteins involved in selective autophagy.
In order to determine whether galectins are required for autophagy protein recruitment, macrophages deficient in galectins-3, -8 or -9 were generated. Galectin-8 deficient macrophages had reduced LC3 recruitment to M. tuberculosis, as well as an increase in bacterial replication, suggesting that galectin-8 promotes M. tuberculosis targeting to autophagic machinery in macrophages.
The next key finding is that galectin-8 can interact with ubiquitin and a known selective autophagy receptor TAX1BP1. This led the authors to hypothesise that galectin-8 binding with TAX1BP1 increases selective autophagy at sites of membrane damage. TAX1BP was next shown to co-localise with galectin-8-positive M. tuberculosis in macrophages, but this did not occur using an M. tuberculosis strain which was unable to cause membrane damage. TAX1BP1 recruitment to M. tuberculosis was also reduced in cells deficient in galectin-8. Finally, galectin-8 was overexpressed in macrophages, leading to an increase in LC3 recruitment to M. tuberculosis. Together these data suggest that galectin-8 promotes selective autophagy adaptor recruitment to damaged M. tuberculosis phagosomes helping to control infection.
Why we chose this preprint:
We have chosen this study because this preprint examines host-pathogen interactions leading to potential therapeutic targets for treatment of tuberculosis, similar to this study recently highlighted. This study points to host cell danger sensors—galectins—as potential therapeutic targets to promote the clearance of M. tuberculosis. In addition, the imaging of multiple protein interactions with M. tuberculosis in macrophages used in this study clearly demonstrated the extent of protein recruitment.
Questions to the authors:
1. Do you think galectins-3 and/or -9 may play alternative roles in promotion of selective autophagy or alternative macrophage antimicrobial defence mechanisms?
2. Would you expect to find a similar role of galectin-8 and selective autophagy in tuberculosis infection in other phagocytes?
3. Do you think galectin-8 and selective autophagy may be important for control of other intracellular pathogens which gain access to the cytosol?
References
- World Health Organisation. Global tuberculosis report 2019. World Health Organization; 2019.
- Farré J-C, Subramani S. Mechanistic insights into selective autophagy pathways: lessons from yeast. Nat Rev Mol Cell Biol. 2016 Sep 6;17(9):537–52.
- Watson RO, Manzanillo PS, Cox JS. Extracellular M. tuberculosis DNA targets bacteria for autophagy by activating the host DNA-sensing pathway. Cell. 2012 Aug 17;150(4):803–15.
- Thurston TLM, Wandel MP, Muhlinen N von, Foeglein Á, Randow F. Galectin-8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature. 482(7385):414.
doi: https://doi.org/10.1242/prelights.23220
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the microbiology category:
Gut microbiome changes over the course of multiple sclerosis differentially influence autoimmune neuroinflammation
Carole Djagang et al.
Human Assembloid Model of Emergent Neurotropic Enteroviruses
Mitchell Sarmie
Cryo-EM reveals multiple mechanisms of ribosome inhibition by doxycycline
Leonie Brüne
preLists in the microbiology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
BioMalPar XVI: Biology and Pathology of the Malaria Parasite
[under construction] Preprints presented at the (fully virtual) EMBL BioMalPar XVI, 17-18 May 2020 #emblmalaria
| List by | Dey Lab, Samantha Seah |
1
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |











 (No Ratings Yet)
(No Ratings Yet)