A copper chaperone-mimetic polytherapy for SOD1-associated amyotrophic lateral sclerosis
Posted on: 9 April 2021
Preprint posted on 23 February 2021
2 is better than 1: the synergistic effect of ebselen and CuATSM for ALS treatment
Selected by Utrecht Protein Folding and AssemblyCategories: biochemistry
Written by: Sanne van Falier, Laurens Geene, Janny Liebregts, and Lucas Siero
Background
Proteins need to fold correctly to exert their function. Misfolded proteins can lead to aggregate formation, which occurs in a variety of diseases. One such disease, amyotrophic lateral sclerosis (ALS), is a severe neurological disease which causes loss of voluntary muscle movement. Certain genetic ALS variations are associated with the misfolding of mutant forms of the protein superoxide dismutase (SOD1). SOD1 folding is stabilized through the insertion of both a Zn and Cu ion in the metal binding region (MBR). Mutations in this region (MBR mutants) lead to impeded ion insertion and enzymatic activity. Correct SOD1 folding also relies on the formation of a disulfide bridge between Cys57-Cys146, and aberrant folding without the disulfide bridge inhibits the dimerization of SOD1 needed for full enzymatic function. Wild-type like (WTL) mutants still show enzymatic activity, but make SOD1 more prone to aggregation.
Normally, the CCS chaperone facilitates copper insertion and disulfide bond formation, but it is unable to fully do so in mutated SOD1. In this preprint, the authors test two small molecules to recapitulate CCS function in a combination treatment: CuATSM, a drug already in clinical trials which facilitates copper ion insertion into SOD1 [1], and ebselen, which promotes the formation of the Cys57-Cys146 disulfide bridge [2]. While both drugs are known to independently rescue disease related phenotypes in mice [1,2], it is yet unknown whether they can be applied in a synergistic way for ALS treatment.
Results
To be able to study the effect of ebselen on SOD1 aggregation, the authors express EGFP-tagged SOD1 in NSC-34 motor neuron-like cells, resulting in the formation of observable inclusions. To identify cells containing inclusions, the authors optimize an analytical pipeline using CellProfiller software. With the use of machine learning, they find optimal parameters to detect inclusions at an extremely high accuracy.
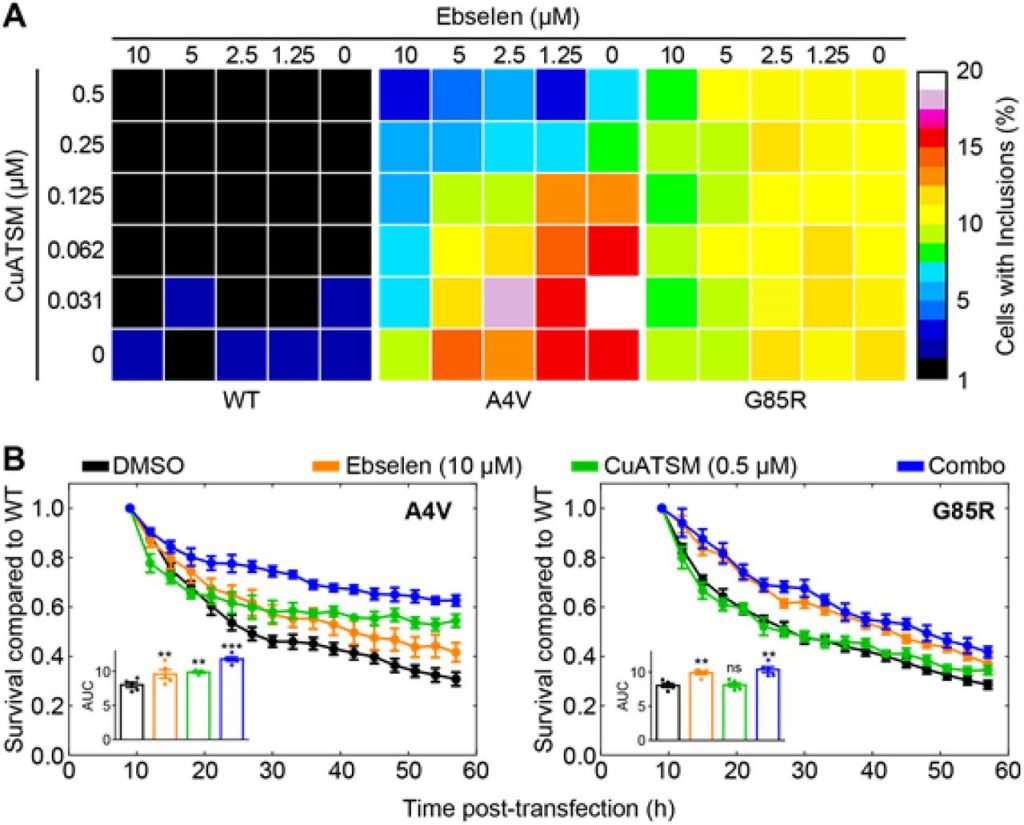
Using this pipeline, they test the effect of ebselen on the inclusion formation of multiple SOD1 mutants, including WTL (A4V) and MBR (G85R) mutants. Treatment with ebselen significantly reduces the formation of inclusions, likely by facilitating the formation of the disulfide bridge. To confirm this, they test the effect of ebselen on a number of mutants including G127X, which deletes residues 127-153. This region contains Cys146 involved in the formation of the disulfide bond. Ebselen has no effect on preventing inclusion formation of SOD1 G127X, but does have a dose dependent effect on other mutants, which can form the disulfide bridge. This finding supports the notion that ebselen facilitates the formation of the disulfide bridge.
Since ebselen cannot take over the copper insertion function, a second small molecule is needed to perform this action. In previous research, CuATSM was found to be capable of delivering copper to SOD1 in cells as well as in mice [1]. A checkerboard analysis using both molecules and WT SOD1 and WTL and MBR mutants shows that the WTL mutant responds to the polytherapy, while the MBR mutant only responds to the ebselen monotherapy. The addition of ebselen decreases the required CuATSM concentration for A4V by a factor 3, while the addition of CuATSM decreases the required amount of ebselen by a factor 10. These results indicate a synergetic drug interaction.
Native SOD1 is active as a dimer. Mutations in the SOD1 gene can negatively affect dimerization and therefore decrease enzymatic activity even further. Treatment with ebselen, CuATSM or the combination therapy rescues this phenotype by increasing dimerization, most prominently for the G93A mutant (a WTL mutation). Furthermore, all mutants show an increase in enzymatic activity after treatment with CuATSM or the combination therapy, but not with ebselen on its own. Polytherapy has the strongest effect for V148G, rescuing the enzymatic activity more than either drug separately.
Altogether, the authors conclude that CuATSM and ebselen can both be used to support correct folding of SOD1 by acting as a CCS mimetic, promoting copper binding and disulfide formation, respectively. The combination of these two drugs can function in a synergistic way for certain SOD1 mutants, which can be exploited for personalised medicine by mutation-specific treatment.
Why we chose this preprint
ALS is a terrible disease with no current treatment. It is important to find therapies that can combat specific ALS mutations, and the application of multidrug therapy can contribute to this. Combination therapy is commonly used in the treatment of cancer, but has never been exploited for ALS. The polydrug therapy tested in this preprint is especially interesting because for certain mutations it not only prevents aggregation, but also partly restores the enzymatic activity of SOD1.
We are also amazed by the accuracy of the optimized machine learning algorithm. By finding the right parameters, this algorithm can be used to screen a large number of cells for the presence of inclusions, which can be applied to find further drugs able to combat ALS and other diseases associated with inclusion formation.
Questions to the authors
Since ebselen was originally designed for hearing loss and therefore aimed at a different target protein, would it be possible to modify the drug making it even more specific for SOD1?
Previous research into ebselen showed it to be less potent at facilitating SOD1 maturation, while a much higher concentration was used (200 µM) [2]. How do you think this is possible?
References
1) Roberts BR, Lim NKH, McAllum EJ, Donnelly PS, Hare DJ, Doble PA, et al. Oral Treatment with CuII(atsm) Increases Mutant SOD1 In Vivo but Protects Motor Neurons and Improves the Phenotype of a Transgenic Mouse Model of Amyotrophic Lateral Sclerosis. Journal of Neuroscience. 2014. pp. 8021–8031. doi:10.1523/jneurosci.4196-13.2014
2) Capper MJ, Wright GSA, Barbieri L, Luchinat E, Mercatelli E, McAlary L, et al. The cysteine reactive small molecule ebselen facilitates effective SOD1 maturation. Nat Commun. 2018;9: 1693.
doi: Pending
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the biochemistry category:
UFMylation of Pyruvate Dehydrogenase Regulates Mitochondrial Metabolism
Hannah Pletcher
Liquid-Liquid Phase Separation-mediated formation of amyloid fibrils from DcpS scavenger enzymes
Joao Gabriel, Marcus Oliveira
4,5-dihydroxyhexanoic acid is a robust circulating and urine marker of mitochondrial disease and its severity
Hannah Pletcher
preLists in the biochemistry category:
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
Peer Review in Biomedical Sciences
Communication of scientific knowledge has changed dramatically in recent decades and the public perception of scientific discoveries depends on the peer review process of articles published in scientific journals. Preprints are key vehicles for the dissemination of scientific discoveries, but they are still not properly recognized by the scientific community since peer review is very limited. On the other hand, peer review is very heterogeneous and a fundamental aspect to improve it is to train young scientists on how to think critically and how to evaluate scientific knowledge in a professional way. Thus, this course aims to: i) train students on how to perform peer review of scientific manuscripts in a professional manner; ii) develop students' critical thinking; iii) contribute to the appreciation of preprints as important vehicles for the dissemination of scientific knowledge without restrictions; iv) contribute to the development of students' curricula, as their opinions will be published and indexed on the preLights platform. The evaluations will be based on qualitative analyses of the oral presentations of preprints in the field of biomedical sciences deposited in the bioRxiv server, of the critical reports written by the students, as well as of the participation of the students during the preprints discussions.
| List by | Marcus Oliveira et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
20th “Genetics Workshops in Hungary”, Szeged (25th, September)
In this annual conference, Hungarian geneticists, biochemists and biotechnologists presented their works. Link: http://group.szbk.u-szeged.hu/minikonf/archive/prg2021.pdf
| List by | Nándor Lipták |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Cellular metabolism
A curated list of preprints related to cellular metabolism at Biorxiv by Pablo Ranea Robles from the Prelights community. Special interest on lipid metabolism, peroxisomes and mitochondria.
| List by | Pablo Ranea Robles |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |











 (No Ratings Yet)
(No Ratings Yet)