LINE elements are a reservoir of regulatory potential in mammalian genomes
Posted on: 22 June 2020
Preprint posted on 31 May 2020
Dynamic turnover of tissue-specific and tissue-shared regulatory elements during mammalian evolution
Selected by Sergio MencheroCategories: evolutionary biology, genomics
In an organism, different gene expression programs are activated in specific tissues and cell types to ensure particular developmental features and cellular functions. The control of those programs is driven by regulatory elements that enhance, repress, isolate or maintain gene expression in a spatial and temporal manner. Although many genes and specific expression programs are widely conserved among different phyla, the regulatory elements controlling those genes seem to have evolved more quickly (Villar et al. 2015). In this work, the authors characterise regulatory elements in four tissues of ten mammals that cover 160 million years of divergence in order to understand the principles underlying the evolution of tissue-specific and tissue-shared regulatory domains.
Scientists have classified regulatory elements according to their activity based on the presence of specific histone modifications. However, the same regulatory domain can have different activities depending on the cell type, the developmental timing and the species (Dao et al. 2017, Carelli et al. 2018). By means of ChIP-seq, the authors mapped three histone modifications whose combination is associated with specific regulatory activity. They focused on the identification of active promoters (characterised by the presence of H3K4me3 and H3K27ac), active enhancers (H3K4me1 and H3K27ac) and primed enhancers (H3K4me1) in four adult tissues: liver, muscle, brain and testis. In parallel, they performed RNA-seq in the same tissues to match transcriptional activity to the presence of the regulatory elements.
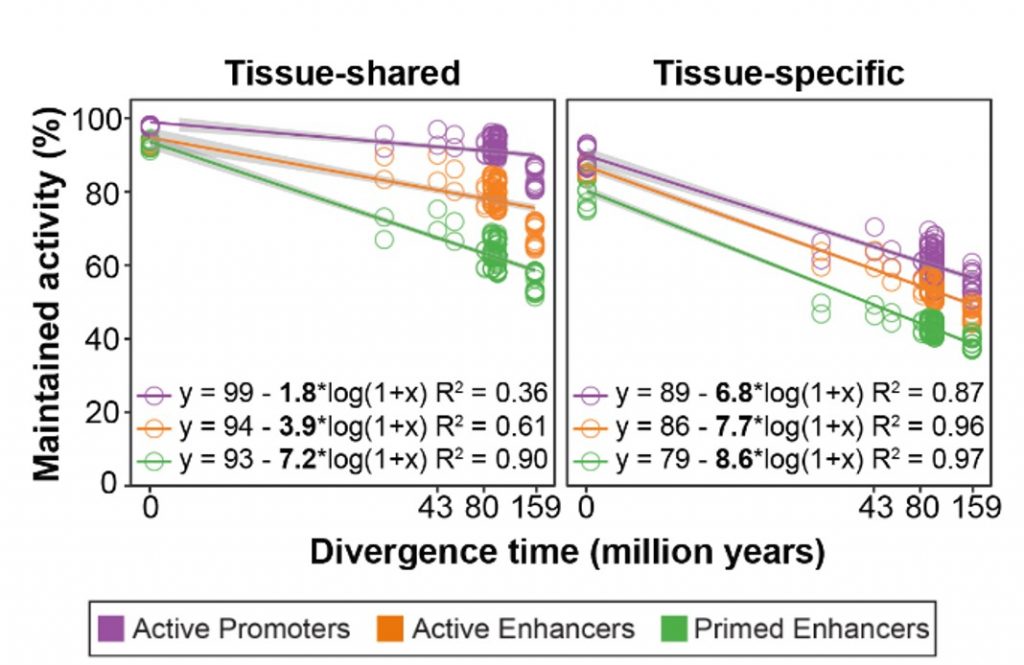
In order to understand the evolutionary stability of the regulatory regions, the authors quantified their evolutionary rates by calculating the fraction of enhancers and promoters that were maintained between pairs of species. The authors found that tissue-shared regulatory regions were more conserved (lower evolutionary rate) than their tissue-specific counterparts. Comparing the different types of regulatory regions, the authors demonstrated that primed enhancers have the highest evolutionary rate while active promoters are the most conserved (Figure 1). Although that is the general trend, there are some differences among tissues. For instance, in testis, active promoters have a higher evolutionary rate than enhancers. Also, brain-specific regulatory regions evolved the most slowly. Remarkably, these particular features correlate with previously reported gene expression evolutionary rates in which small changes in the brain and accelerated changes in testis were described (Cardoso-Moreira et al. 2019).
The authors then focused on the evolutionary turnover of the identified regulatory regions. They defined intra-species dynamic regulatory regions as the regions that have different histone modification signatures across tissues of the same species, and between-species dynamic regulatory regions when the changes in histone modification signatures occurred between different species (in pairwise comparisons). Intra-species dynamic regulatory regions were relatively rare (7-11% of regions were characterised), and the ones identified were not maintained as dynamic regions in other species. In contrast, promoter-enhancer switching were much more common between species. Particularly high was the turnover between active and primed enhancers (44% of active enhancers aligned to a prime enhancer in another species). Taking into account the evolutionary distance, the authors showed that enhancers are more likely to evolve into active promoters than the reverse (six times more!). Nicely, those enhancer-to-promoter switches were correlated with higher transcriptional activity of the associated gene.
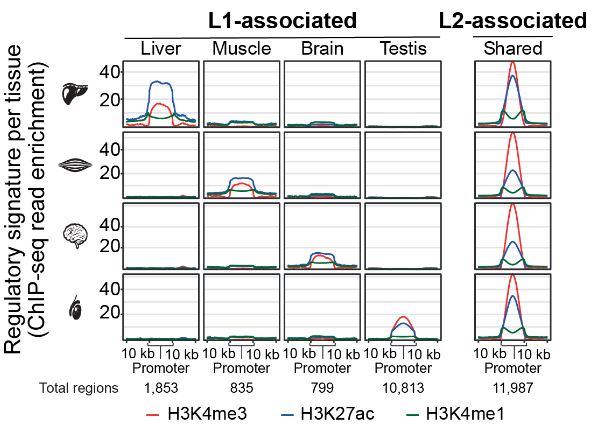
Finally, the authors investigate whether specific classes of transposable elements contribute to the evolution of regulatory elements. They compared the enrichment of annotated transposable elements in the identified tissue-shared and tissue-specific regulatory elements. Interestingly, tissue-specific regulatory elements were enriched in LINE L1 family of transposons while tissue-shared elements were enriched in transposons of the LINE L2 family (Figure 2). When comparing stable active enhancers versus evolutionary dynamic elements, the authors showed that stable enhancers were enriched in the LINE L1 family while dynamic elements were enriched with in LINE L2 family of transposons. This suggests that LINE L2 transposons may provide a more versatile potential for transcriptional regulation.
Why I chose this preprint and why I think this work moves the field forward:
The control of transcription mediated by the activity of regulatory elements has always been close to my own research directly or indirectly (by the work of some colleagues). The identification of regulatory elements is always complicated and we have to rely on the presence of histone marks or reporter activity, which is never 100% accurate. Despite all the variability and exceptions, researchers have managed to improve our understanding of this complex but robust system of gene regulation.
I find particularly fascinating the evolutionary perspective of the regulatory regions. Beautiful works from past years have shown how enhancers evolved from ancestral DNA exaptation (Villar et al. 2015) or how regulatory landscapes change their complexity during evolution to deal with whole-genome duplication and specialization events (Marlétaz et al. 2018). In this work, the characterisation of regulatory elements across ten different mammals and four different tissues allowed the authors to provide a wide view of the evolution of dynamic regulatory landscapes. Due to the complexity and variability of regulatory elements when studied one by one, it is important to first have a global picture of what are the behaviours that mostly underlie enhancer or promoter evolution. All these recent works, including this preprint, are important landmarks for future studies where more particular cases of gene and regulatory landscape evolution can be addressed.
Questions to the authors
- The authors identify regulatory elements based on the presence of specific histone marks for all the species. Have the authors studied if the degree of sequence conservation correlates with regulatory elements being more stable or dynamic?
- The authors have shown how turnover of regulatory elements affected the transcriptional activity of their associated genes. Do the authors think the evolution of genes can also affect the activity of regulatory elements? If a gene is not conserved in a species, due to exon erosion or transposon insertion for instance, can the associated regulatory elements be maintained or would they also disappear?
References
Carelli et al. Repurposing of promoters and enhancers during mammalian evolution. Nat Commun 2018 Oct 4;9(1):4066.
Cardoso-Moreira et al. Gene expression across mammalian organ development. Nature 2019 Jul;571(7766):505-509.
Dao et al. Genome-wide characterization of mammalian promoters with distal enhancer functions. Nat Genet 2017 Jul;49(7):1073-1081
Marlétaz et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 2018 Dec;564(7734):64-70.
Villar et al. Enhancer evolution across 20 mammalian species. Cell 2015 Jan 29;160(3):554-66
doi: https://doi.org/10.1242/prelights.22018
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the evolutionary biology category:
Heatwave winners and losers: cryptic coral holobionts differ in thermal tolerance
Maitri Manjunath
No global collapse of food webs across the Permian–Triassic Mass Extinction
Theodora Stougiannou
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
Also in the genomics category:
Temporal degradation of PRC2 uncovers specific developmental dependencies
María Mariner-Faulí
Gut microbiome changes over the course of multiple sclerosis differentially influence autoimmune neuroinflammation
Carole Djagang et al.
Microbial Feast or Famine: dietary carbohydrate composition and gut microbiota metabolic function
Jasmine Talevi
preLists in the evolutionary biology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
EMBO | EMBL Symposium: The organism and its environment
This preList contains preprints discussed during the 'EMBO | EMBL Symposium: The organism and its environment', organised at EMBL Heidelberg, Germany (May 2023).
| List by | Girish Kale |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
COVID-19 / SARS-CoV-2 preprints
List of important preprints dealing with the ongoing coronavirus outbreak. See http://covidpreprints.com for additional resources and timeline, and https://connect.biorxiv.org/relate/content/181 for full list of bioRxiv and medRxiv preprints on this topic
| List by | Dey Lab, Zhang-He Goh |
1
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |
Also in the genomics category:
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
March in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) cancer biology 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics and genomics 6) other
| List by | Girish Kale et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
Early 2025 preprints – the genetics & genomics edition
In this community-driven preList, a group of preLighters, with expertise in different areas of genetics and genomics have worked together to create this preprint reading list. Categories include: 1) bioinformatics 2) epigenetics 3) gene regulation 4) genomics 5) transcriptomics
| List by | Chee Kiang Ewe et al. |
End-of-year preprints – the genetics & genomics edition
In this community-driven preList, a group of preLighters, with expertise in different areas of genetics and genomics have worked together to create this preprint reading list. Categories include: 1) genomics 2) bioinformatics 3) gene regulation 4) epigenetics
| List by | Chee Kiang Ewe et al. |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Semmelweis Symposium 2022: 40th anniversary of international medical education at Semmelweis University
This preList contains preprints discussed during the 'Semmelweis Symposium 2022' (7-9 November), organised around the 40th anniversary of international medical education at Semmelweis University covering a wide range of topics.
| List by | Nándor Lipták |
20th “Genetics Workshops in Hungary”, Szeged (25th, September)
In this annual conference, Hungarian geneticists, biochemists and biotechnologists presented their works. Link: http://group.szbk.u-szeged.hu/minikonf/archive/prg2021.pdf
| List by | Nándor Lipták |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
Zebrafish immunology
A compilation of cutting-edge research that uses the zebrafish as a model system to elucidate novel immunological mechanisms in health and disease.
| List by | Shikha Nayar |











 (No Ratings Yet)
(No Ratings Yet)