NeuroMechFly, a neuromechanical model of adult Drosophila melanogaster
Posted on: 23 June 2021
Preprint posted on 17 May 2021
Article now published in Nature Methods at http://dx.doi.org/10.1038/s41592-021-01226-z
Categories: bioengineering, neuroscience
Background
Movement is complicated. Body movement, be it walking, swimming, or flying, is an outcome of the biomechanics of the limbs that propel the animal and the neural circuits that exert control over limb movements. Because animal behaviors typically occur in response to environmental cues, it is easy to forget the importance of biomechanics and assume that neural circuits control all locomotion.
The active control of movement via muscles depends on the timing of muscle activation, based on which muscles can function as motors, brakes, or even as struts and springs[1]. Even the sensory input to the nervous system is affected by the biomechanics of the limb. Proprioceptors, which are sensors that provide the nervous system with real-time information of body position, velocity, etc., are strongly influenced by their position on the body and the mechanics of body movement[2]. All these suggest that biomechanics have a critical influence on the computations that need to be performed by neural circuits and the type of control required for locomotion.
One way to understand this interplay is to build neuromechanical models that integrate both biomechanics and neural control. However, this is easier said than done, as these models sit at the interface of biology, mechanics, and neuroscience. Traditionally, such models are made for one type of locomotion, like for swimming in fish and for walking in stick insects. It is difficult to build neuromechanical models of the whole animal that can be potentially used to simulate the whole behavioral repertoire of an animal.
In this preprint, Ríos et al. develop the first data-driven neuromechanical model of the fruit fly, Drosophila melanogaster, a widely studied model organism with complex behaviors and a mapped neural connectome. Using this model, dubbed NeuroMechFly, the authors show how such models can be used to investigate the interplay between biomechanics and neural control that gives rise to behavior.
Key Findings
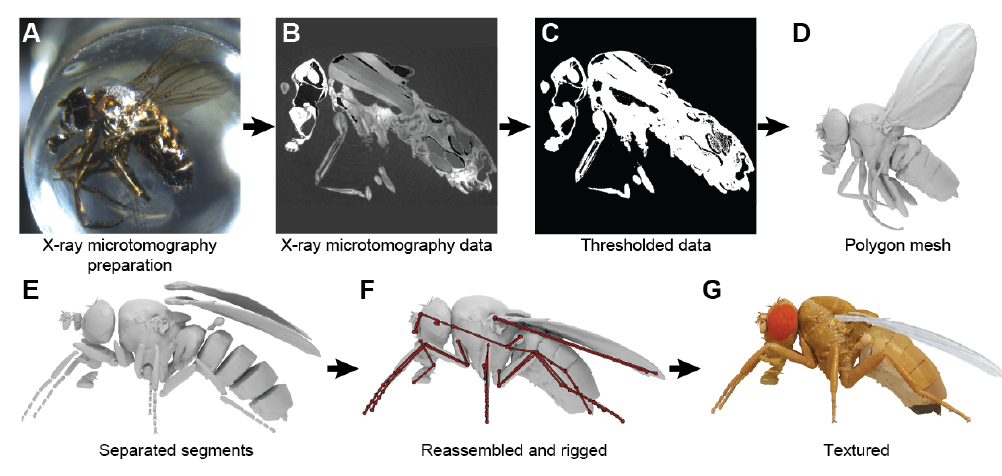
- The authors used x-ray microtomography to scan an adult fruit fly (Fig. 1). Using this data, they created 3D meshes for the exoskeleton of the fly, and segments for individual parts such as the wings, legs, antennae, and halteres. Joints were added between the segments, which allowed the body parts to be moved (Video 1). This model formed the base over which joint biomechanics, and active control via muscles and neural circuits were built.
- Next, the authors focused on walking and grooming behaviors in insects. To each leg joint in the neuromechanical model they added known degrees of freedom (DoF), which is essentially the axis along which the joint can move. By capturing 3D movement data from live flies walking and grooming, the authors could now compare the performance of the neuromechanical model with data from live flies. Surprisingly, they found a clear difference – the model based on known DoFs were not sufficient to explain all the poses seen in walking/grooming flies. To get the model to behave like a live fly, they needed to add an extra DoF to the trochanter-femur joint, one of the joints of the insect leg. This joint was earlier thought to be fused in fruit flies and act like a rigid structure. The use of the model to assess the current understanding of joint movement provided a prediction that could be experimentally tested, i.e., is the trochanter-femur joint capable of movement along one axis?
- Using a physics-based simulation environment, the authors explored a way to extract contact forces and joint torques from the model. As a first step, the neuromechanical model was made to move in a fly-like manner by adding a controller to each of the joint legs and tuning them using movement data from live flies. By simulating the model fly walking, the authors were then able to measure the contact forces on the fly’s legs and torques on individual leg joints (Video 1). Although the forces and torques obtained from these simulations still require experimental validation, they serve as a proofs-of-concept of the use of such (calibrated) models to estimate hard-to-measure quantities.
- Finally, to illustrate the use of such models in understanding neuromuscular dynamics, they authors added in muscles and neurons to the neuromechanical model. For these simulations, they restricted the model by only adding muscles to the joints that were critical for walking. Each joint consisted of two muscles – a flexor and an extensor – and a neural controller that controlled these two muscles. Using this, they were able to find control parameters that produced tripod gaits and look at the dynamics of neural control and muscle activation that gave rise to the gait (Video 2). This too served as a proof-of-concept of using such models to glean the workings of neural circuits that can control these movements.
Why I think this work is important
I feel that a complete neuromechanical model, such as the one described in this paper, enable scientists who work on different parts of the fly to pool their data in a scalable manner. For instance, it potentially allows two groups that work on walking flies and flying flies to pool their data and obtain a model that can do both. In a sense, it attempts to solve the chaos in the brickyard problem[3], at least for the interface of biomechanics and the neural control of movement.
Future directions / Questions to authors
- MicroCT data also provides information about the muscles, their attachment points, etc. and about proprioceptors and mechanosensors embedded on the insect cuticle. Incorporating this would constrain inputs to and outputs from the neural controllers.
- Do you have guesses on how easy/hard it would be to incorporate this information into the biomechanical model?
- Getting information about muscles and proprioceptors would require microCT scans with different resolutions (a guess – microCT novice here). If so, are there challenges in incorporating data with different resolutions (voxel sizes maybe)?
- Validation of predictions and calibration of the neuromechanical model is a critical step before the model can be used in place of experimental measurements. However, I presume this is going to be incredibly challenging. If possible, could you share your plan on how you will go about it?
- This neuromechanical model is a brilliant resource that can be scaled up and developed together as a community. The code is open source, but often this is not sufficient to develop a community that will use this resource. Building a platform around it usually helps. Do you plan to build one that makes it easy to generate models and add features?
References
[1] Dickinson, M. H., Farley, C. T., Full, R. J., Koehl, M. A. R., Kram, R., & Lehman, S. (2000). How animals move: an integrative view. Science, 288(5463), 100-106.
[2] Sane, S. P., & McHenry, M. J. (2009). The biomechanics of sensory organs. Integrative and Comparative Biology, 49(6), i8-i23.
[3] Check out the comic adaptation and the associated paper: Forscher, B. K. (1963). Chaos in the brickyard. Science, 142(3590), 339-339
doi: https://doi.org/10.1242/prelights.29752
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the bioengineering category:
Detergent-Triggered Membrane Remodelling Monitored via Intramembrane Fluorescence De-Quenching
Cyntia Alves Conceição, Marcus Oliveira
A Novel Chimeric Antigen Receptor (CAR) - Strategy to Target EGFRVIII-Mutated Glioblastoma Cells via Macrophages
Dina Kabbara
Human pluripotent stem cell-derived macrophages modify development of human kidney organoids
Theodora Stougiannou
Also in the neuroscience category:
Behavioral characteristics of an extremely old rhesus macaque in a zoo: Dementia-like symptoms and implications for quality of life of geriatric animals
Stefan Friedrich Wirth
EBV reprograms autoreactive anti-CNS B cells as antigen presenting cells in multiple sclerosis
Léa Bastien et al.
The Endocannabinoid System’s Contribution to Placebo Analgesia
Thomas Nicodemo Arrieta et al.
preLists in the bioengineering category:
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Advances in microscopy
This preList highlights exciting unpublished preprint articles describing advances in microscopy with a focus on light-sheet microscopy.
| List by | Stephan Daetwyler |
Also in the neuroscience category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |











 (No Ratings Yet)
(No Ratings Yet)