A Single Laccase Acts as a Key Component of Environmental Sensing in a Broad Host Range Fungal Pathogen
Posted on: 16 May 2023
Preprint posted on 12 January 2023
Article now published in Communications Biology at http://dx.doi.org/10.1038/s42003-024-06034-7
Roses are red, Violets are blue, S. sclerotiorum is a rot-causing fungal plant pathogen, And its virulence depends on Sslac2! preLight Authors: Annika Schulz & Shamiul Rasel
Selected by UofA IMB565Categories: genetics, microbiology, molecular biology, plant biology
preLight Authors: Annika Schulz & Shamiul Rasel
Background:
Laccases are enzymes that play a crucial role in mediating the interaction between lignin production and degradation in plants and fungi, which is critical to carbon cycling and maintaining healthy soils. In ascomycetes (fungi that belong to the phylum Ascomycota and generally produce sexual spores in sacs (1, 2)), laccases are typically associated with melanin or pigment deposition on fungal tissue, and most laccase knockout mutants within the phylum exhibit reduced pigmentation. However, the effect of laccase knockouts on fungal development and growth varies widely, possibly due to the expanded repertoire of laccases and functional redundancy. Specific laccase genes have been noted to play a role in virulence on host plants, such as in Colletotrichum gloeosporioides, Colletotrichum orbiculare, and Setosphaeria turcica, where individual laccase gene knockouts resulted in reduced virulence. Because laccases are often associated with pigmentation, the lack of melanin/pigment deposition in the knockout is assumed to be responsible for these observed phenotypes. However, no causal relationship has been established.
Sclerotinia sclerotiorum is a well-known fungal plant pathogen that causes rot/lesions in various plant parts. It can attack a wide range of plant species, including important crops like oilseed rape, soybean, and other vegetables (3), causing serious economic losses globally (4). Plant cell wall-degrading enzymes released by S. sclerotiorum soften, hydrolyze, and degrade plant cells (3, 4). S. sclerotiorum’s oxalic acid disrupts host cell metabolism after destroying the plant cell wall (4). In S. sclerotiorum, previous studies found that knocking out genes involved in melanin biosynthesis did not affect virulence. Nonetheless, little is known about the functions of laccases as virulence factors.
In this preprint, a laccase gene called Sslac2 was identified in S. sclerotiorum and found to be highly upregulated during infection of soybean plants. This study discusses the role of Sslac2 as a global regulator of environmental sensing, affecting every stage of development, infection, and response to environmental cues, which is unlike the laccase gene in a closely related species. The preprint also highlights the potential for targeting fungal laccases to increase resistance in host plants and considers the broader context of laccases within fungal plant pathogen biology.
Key findings
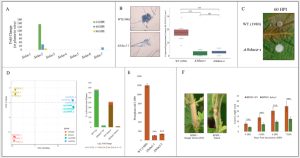
Figure 1. Composition of several figures from Westrick et al. (A) Sslac2 is highly upregulated during early stages of infection. (B) Mutants lacking Sslac2 produce malformed appressoria. (C) Mutants lacking Sslac2 are essentially unable to cause disease. (D) WT shows a transcriptional shift in response to GrSt. ΔSslac2 is unresponsive to GrSt and shows a lack of differential gene expression compared to WT. (E) ΔSslac2’s hyphal cell wall is resistant to protoplasting upon treatment with cell wall degrading enzymes. (F) Silencing Sslac2 reduces the size of lesions in infected plants. The reddening around the lesion indicates a resistance response from the host plant.
Sslac2 is upregulated during infection and implicated in fungal growth
Fungal laccases have been previously implicated in plant pathogenicity, which lead Westrick and colleagues to investigate the seven laccase-encoding genes in S. sclerotiorum. Comparing the regulation of these genes during infection (when grown in planta vs. in vitro), Sslac2 was the only laccase markedly upregulated (Fig. 1A). This upregulation likely occurs when S. sclerotiorum contacts the plant.
The Sslac2 mutant S. sclerotiorum strain (ΔSslac2) was unable to produce sclerotia (masses of hyphae used for survival (5)) and had significantly decreased laccase activity that was not rescuable with other laccase homologues. These results indicate that when S. sclerotiorum is present on solid surfaces (i.e., plants), it mainly uses Sslac2 for growth and development.
ΔSslac2 mutants are essentially non-pathogenic
ΔSslac2 mutants had extremely malformed appressoria (structures formed by the fungus for the purpose of penetrating the host), resulting in difficulty perforating and invading plant tissue. Also, they produced less oxalic acid, which is important for host tissue acidification, cell death inducement, and host defense subversion (Fig. 1B). These attributes, along with additional suspected defects, rendered ΔSslac2 essentially non-pathogenic (Fig. 1C). The authors speculated that ΔSslac2’s virulence defects were likely stemming from other issues, like a hindered ability to respond to environmental stress (induced by the host). In order to analyze this, they compared the gene expression profiles of ΔSslac2 and the WT. When in the presence of GrSt (soybean green stem extract), the WT’s gene expression profile – unlike the mutant’s – showed a major transcriptional shift, representing its response to the proteins, carbohydrates, and chemical signals from the plant (Fig. 1D). Mutants ΔSslac1 and 2 had significantly fewer differentially expressed genes compared to the WT and were essentially non-responsive to GrSt (Fig. 1D).
ΔSslac2 mutants cannot properly sense and respond to host-induced environmental stress
Westrick and colleagues next sought to establish the factors contributing to ΔSslac’s environmental sensing defect. Because ascomycete fungi laccases are generally implicated in cellular detoxification and cell wall remodeling, they analyzed the mutant’s sensitivity to different plant antifungal compounds and cell wall stressors. When S. sclerotiorum was treated with toxic plant-derived compounds, the ΔSslac mutants were significantly less resistant than the WT. When treated with cell wall stressors, the ΔSslac mutants were again significantly less resistant. The authors next investigated the basis of ΔSslac2’s increased susceptibility to cell wall stressors. They found that ΔSslac2 was significantly more resistant to protoplasting by cell wall degrading enzymes (Fig. 1E), had structural differences in hyphae/hyphal growth, including decreased hyphal hydrophobicity, and had textural differences in their extracellular matrix (ECM).
Westrick and colleagues observed decreased radial growth for ΔSslac2 grown on PDA (potato dextrose agar) plates but no decrease in mass in liquid media, indicating a specific radial growth defect. They also observed differences in hyphae for ΔSslac2 cultures. ΔSslac2 had increased aerial hyphae that, unlike the WT, grew upwards over the plate dividers and downward into the agar. Because the WT would instead become dormant and begin building survival structures (like sclerotia), the authors hypothesized that Sslac2 may play a role in sensing environmental signals for dormancy or sclerotia production. In order to assess this, they needed to determine whether the differences in ΔSslac2’s radial growth defect and hyphae (agar penetration) were due to issues with directional growth or with responding to environmental dormancy signals. Unlike when grown on PDA, ΔSslac2 grown on cellophane did not have decreased radial growth. This result, taken into account with the lack of sclerotia production, suggests that ΔSslac2 cannot sense the environmental signals for dormancy and instead grows randomly. Therefore, Sslac2 may be implicated in hyphal thigmotropism and differentiation.
Host-induced gene silencing of Sslac2 reduces the virulence of S. sclerotiorum
Silencing Sslac2 via VIGS (vector-induced gene silencing) decreased the sizes of the lesions formed on plants by S. sclerotiorum. There was also visible reddening around the lesions on the soybean plants’ stems that were inoculated with a Sslac2 silencing vector, which indicated a host defense response from the plant. These results suggest that silencing Sslac2 reduces the virulence of S. sclerotiorum and allows the host to spend more time mounting a defense (Fig. 1F). Because VIGS was successful, targeting Sslac2 via HIGS (host-induced gene silencing) might be a viable method of disease control.
Summary
Overall, this preprint determined that Sslac2, a laccase present in the fungal plant pathogen S. sclerotiorum, is a key player in virulence. It contributes to fungal growth and development and environmental signal sensing, allowing the pathogen to successfully infect its host. Because of these critical roles in virulence and the fact that silencing Sslac2 resulted in decreased virulence, Sslac2 may be an attractive target for fungal disease control in plants.
What we like:
This study employs a combination of genetic and biochemical approaches to investigate the function of a particular laccase enzyme. The methods used are well-established in the field of fungal biology, which adds credibility to the study, and the conclusions drawn are well-supported by the results. This study is especially interesting, as little research has been done to establish the roles of secreted fungal laccases in the pathogenesis of plants. The findings indicate a relatively novel role for Sslac2 in environmental sensing that is important for the virulence of S. sclerotiorum. Understanding the mechanisms that underpin fungal pathogens’ virulence is essential for developing effective strategies to control their spread. Determining the roles of Sslac2 in virulence opens the door to disease treatment via targeting Sslac2, which the authors demonstrate may be feasible using HIGS. Considering S. sclerotiorum infects a broad host range of plants, including many crops, treatment has important implications in agriculture and the field of plant pathology.
Future directions and questions:
The authors state a few goals/possibilities for future work, including determining substrates of Sslac2 and the specific methods through which Sslac2 functions to regulate thigmotropism and environmental signal response. They also plan on using gene silencing to attempt to target Sslac2 and other laccases to treat fungal infection of plants. Another beneficial future direction could include testing laccase deletion mutants on multiple types of plants grown in various conditions–this would help to identify potential varying roles of laccases when subjected to differing conditions.
We wonder what other proteins or pathways may be involved in environmental sensing and how they interact with the laccase-mediated pathway. We also are interested in the potential implications of deleting Sslac2 (and other laccases) on general cellular processes.
References:
(1) M. McConnaughey, (2014). Physical Chemical Properties of Fungi, Reference Module in Biomedical Sciences, Elsevier, ISBN 9780128012383, https://doi.org/10.1016/B978-0-12-801238-3.05231-4.
(2) Nicholas P. Money, (2016). Chapter 1 – Fungal Diversity, Editor(s): Sarah C. Watkinson, Lynne Boddy, Nicholas P. Money, The Fungi (Third Edition), Academic Press, Pages 1-36, ISBN 9780123820341, https://doi.org/10.1016/B978-0-12-382034-1.00001-3.
(3) Daohong Jiang, Yanping Fu, Li Guoqing, Said A. Ghabrial., (2013). Chapter Eight – Viruses of the Plant Pathogenic Fungus Sclerotinia sclerotiorum. Editor(s): Said A. Ghabrial, Advances in Virus Research, Academic Press, Volume 86, Pages 215-248. https://doi.org/10.1016/B978-0-12-394315-6.00008-8.
(4) Yang, C., Li, W., Huang, X., Tang, X., Qin, L., Liu, Y., Xia, Y., Peng, Z., & Xia, S., (2022). SsNEP2 Contributes to the Virulence of Sclerotinia sclerotiorum. Pathogens (Basel, Switzerland), 11(4), 446. https://doi.org/10.3390/pathogens11040446.
(5) Money, Nicholas P., (2023). Chapter 2 – Fungal Cell Biology and Development, Editor(s): Sarah C. Watkinson, Lynne Boddy, Nicholas P. Money, The Fungi (Third Edition), Academic Press, Pages 37-66, ISBN 9780123820341, https://doi.org/10.1016/B978-0-12-382034-1.00002-5.
Reference for the preprint this preLight covers:
Nathaniel M. Westrick, Eddie G. Dominguez, Christina M. Hull, Damon L. Smith, Mehdi Kabbage. A Single Laccase Acts as a Key Component of Environmental Sensing in a Broad Host Range Fungal Pathogen. bioRxiv 2023.01.12.523834, https://doi.org/10.1101/2023.01.12.523834.
doi: Pending
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the genetics category:
Chromatin priming and co-factor availability shape lineage response to the neuronal pioneer factor ASCL1 in pluripotency
Reinier Prosee, Jethro Lundie-Brown
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
Classical enhancers couple cis-regulatory logic with transcriptional condensates and 3D genome architecture
Siddharth Singh
Also in the microbiology category:
Gut microbiome changes over the course of multiple sclerosis differentially influence autoimmune neuroinflammation
Carole Djagang et al.
Human Assembloid Model of Emergent Neurotropic Enteroviruses
Mitchell Sarmie
Cryo-EM reveals multiple mechanisms of ribosome inhibition by doxycycline
Leonie Brüne
Also in the molecular biology category:
Chromatin priming and co-factor availability shape lineage response to the neuronal pioneer factor ASCL1 in pluripotency
Reinier Prosee, Jethro Lundie-Brown
Temporal degradation of PRC2 uncovers specific developmental dependencies
María Mariner-Faulí
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
Also in the plant biology category:
A drought stress-induced MYB transcription factor regulates pavement cell shape in leaves of European aspen (Populus tremula)
Jeny Jose
Actin Counters Geometry to Guide Plant Cell Division
Jeny Jose
The nucleus follows an internal cellular scale during polarized root hair cell development
Jeny Jose
preLists in the genetics category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
March in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) cancer biology 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics and genomics 6) other
| List by | Girish Kale et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
Early 2025 preprints – the genetics & genomics edition
In this community-driven preList, a group of preLighters, with expertise in different areas of genetics and genomics have worked together to create this preprint reading list. Categories include: 1) bioinformatics 2) epigenetics 3) gene regulation 4) genomics 5) transcriptomics
| List by | Chee Kiang Ewe et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
End-of-year preprints – the genetics & genomics edition
In this community-driven preList, a group of preLighters, with expertise in different areas of genetics and genomics have worked together to create this preprint reading list. Categories include: 1) genomics 2) bioinformatics 3) gene regulation 4) epigenetics
| List by | Chee Kiang Ewe et al. |
BSDB/GenSoc Spring Meeting 2024
A list of preprints highlighted at the British Society for Developmental Biology and Genetics Society joint Spring meeting 2024 at Warwick, UK.
| List by | Joyce Yu, Katherine Brown |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
Semmelweis Symposium 2022: 40th anniversary of international medical education at Semmelweis University
This preList contains preprints discussed during the 'Semmelweis Symposium 2022' (7-9 November), organised around the 40th anniversary of international medical education at Semmelweis University covering a wide range of topics.
| List by | Nándor Lipták |
20th “Genetics Workshops in Hungary”, Szeged (25th, September)
In this annual conference, Hungarian geneticists, biochemists and biotechnologists presented their works. Link: http://group.szbk.u-szeged.hu/minikonf/archive/prg2021.pdf
| List by | Nándor Lipták |
2nd Conference of the Visegrád Group Society for Developmental Biology
Preprints from the 2nd Conference of the Visegrád Group Society for Developmental Biology (2-5 September, 2021, Szeged, Hungary)
| List by | Nándor Lipták |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Zebrafish immunology
A compilation of cutting-edge research that uses the zebrafish as a model system to elucidate novel immunological mechanisms in health and disease.
| List by | Shikha Nayar |
Also in the microbiology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
BioMalPar XVI: Biology and Pathology of the Malaria Parasite
[under construction] Preprints presented at the (fully virtual) EMBL BioMalPar XVI, 17-18 May 2020 #emblmalaria
| List by | Dey Lab, Samantha Seah |
1
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |
Also in the molecular biology category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |
Also in the plant biology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |











 (5 votes)
(5 votes)