The Histone H3-H4 Tetramer is a Copper Reductase Enzyme
Posted on: 18 July 2018 , updated on: 3 July 2020
Preprint posted on 19 June 2018
Article now published in Science at https://science.sciencemag.org/content/369/6499/59
New study finds an unprecedented, but ancient, role for histones as enzymes. The histone H3-H4 tetramer is a copper reductase that may have facilitated the emergence of eukaryotes.
Selected by Lauren NevesCategories: molecular biology
Background
2.45 billion years ago, the rise of oxygen-producing bacteria resulted in drastic oxygen accumulation in the Earth’s atmosphere. In addition to organic compounds, such as amino acids, carbohydrates and nucleic acids, a number of transition metals are important for the chemistry of living systems. However, this Great Oxidation Event led to significant changes in abundance and oxidation states of such transition metals that not only altered their usability, but also increased their toxicity.
Copper is a required co-factor for enzymes in a number of biological pathways including oxidative respiration, anti-oxidant defense, melanin production, and tyrosine metabolism. Although the majority of environmental copper is found in the oxidized cupric (Cu2+) form, it is the cuprous (Cu+) form that is transported intracellularly. It is not well understood how this pool of cuprous ions is maintained in the cell.
Interestingly, eukaryotic nucleosomes, the basic structural unit of chromatin, have been found to bind copper and other ions, although the functionality of this binding is unknown. Nucleosomes are composed of DNA wrapped around two copies of each histone proteins H2A, H2B, H3 and H4. Histone proteins are not only important structural elements for DNA, but also have additional regulatory roles. Intriguingly, certain archaeal prokaryotes have nucleosome-like structures despite the fact that they have small genomes, no nucleus and no apparent epigenetic regulation, suggesting that histones may have another function that predates the emergence of eukaryotes.
Attar, Campos, Vogelauer and colleagues have uncovered a surprising, but likely ancient, role for the nucleosome as a copper reductase. They show that the histone H3-H4 tetramer has conserved enzymatic activity that generates usable copper for multiple cellular processes. Given that the evolution of eukaryotes coincided with the Great Oxidation Event, these findings suggest that enzymatic activity of ancient histones may have contributed to the emergence of eukaryotes by providing usable copper to early mitochondria.
Key Findings
Within the assembled nucleosome, a pair of histidine residues (H3H113), one from each apposing H3 histone, help form a pocket capable of binding metal ions. The authors observe that mutation of the histone H3 histidine to asparagine (H3H113N) leads to defects in pathways known to require copper. Yeast cells containing this mutation (H3H113N) require a large excess of copper for proper mitochondrial respiration and lysine biosynthesis. Defects are particularly pronounced upon deletion of a Cu+-ion specific transporter, suggesting that H3H113N mutation decreases intracellular Cu+ levels.
Surprisingly, the authors observe that overall levels of intracellular copper are unaffected in H3H113N cells. Additionally, gene expression and chromatin accessibility are unchanged by H3H113N mutation. Because defects observed in H3H113N cells indicate a loss of cuprous ions, the authors suspected that histone H3 could directly affect copper oxidation. Indeed, in a series of elegant biochemical assays, the authors show that histone H3, when present as a tetramer with H4, has enzymatic activity and catalyzes the reduction of Cu2+ to Cu+ (Figure 4B). Consistent with phenotypic observations, the H3H113N mutation decreases the copper reducing activity of H3-H4 tetramers.
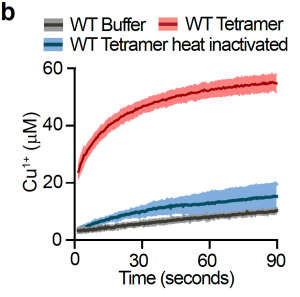
Figure 4B from the preprint: Progress curves of copper reduction by human H3.2-H4 tetramer, heat-inactivated tetramer or buffer.
Both human and yeast tetramers reduce copper, suggesting that catalytic activity is conserved throughout eukaryotes. However, yeast histone H3-H4 tetramers did not reduce copper as well as human H3. It has been shown that a cysteine residue (H3C110), which exists in most species but is lacking in yeast, helps form the metal-binding pocket. The authors find that the presence of H3C110 substantially increases the enzymatic activity of human histone H3-H4 tetramers. In fact, introducing H3C110 in to yeast H3 not only increased copper reduction, but also improved respiratory growth in copper limiting conditions.
Thoughts and future directions
Histones were once thought to be no more than inert structural proteins: packing material for eukaryotic DNA. However, they have proven to have some of the most dynamic roles in the cell. Histones have important roles in DNA replication, gene expression regulation, transcription elongation dynamics, RNA processing, and DNA damage. And now, Attar, Campos, Vogelauer and colleagues have revealed a completely new role that changes the way we think about histones. DNA is wrapped around an enzyme, whose catalytic activity has important consequences for processes throughout the cell. Most excitingly, this role likely predates many of the previously known functions of histones. Reductase activity of early histones likely increased availability of cuprous ions to important biological pathways, which allowed for the rise of more complex organisms and facilitated eukaryogenesis.
Some questions I have:
- Do archaeal histones have copper reductase activity? Evidence of this will strongly support the hypothesis that the enzymatic role of histones predates, and helped facilitate, eukaryogenesis.
- Histone H3 has been shown to bind several metal ions. The authors show that defects in respiration and lysine biosynthesis are specific to copper, but are histones capable of reducing other similar metal ions, such as iron?
- Adamczyk et al. observed that metal binding by histone H3 is not readily reversible. Given that these reduced ions must be available for use outside the nucleus, how are cuprous ions released from the nucleosome?
Further reading
- Adamczyk M, Poznanski J, Kopera E, Bal W. 2007. A zinc-finger like metal binding site in the nucleosome. FEBS Lett 581: 1409-1416.
- Festa RA, Thiele DJ. 2011. Copper: an essential metal in biology. Curr Biol 21: R877-883.
- Kouzarides T. 2007. Chromatin modifications and their function. Cell 128: 693-705.
Sign up to customise the site to your preferences and to receive alerts
Register hereAlso in the molecular biology category:
CENP-B binds hairpin motifs in chromosome arms influencing gene expression
Pierre Caron
Deep Invaginations of Nuclear Envelope Coordinate Spatial Organization of Chromatin in Epithelium
Abby Hein
Ribosome Molecular Aging Shapes Translation Dynamics
Scott Murray Cors
preLists in the molecular biology category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |











 (1 votes)
(1 votes)