Metabolic regulation of species-specific developmental rates
Posted on: 17 September 2021
Preprint posted on 30 August 2021
Article now published in Nature at http://dx.doi.org/10.1038/s41586-022-05574-4
Mouse vs human embryo development - Mouse embryos ace the race! Investigation of metabolic regulation of developmental speed.
Selected by Sundar Naganathan, Julia GrzymkowskiCategories: developmental biology, evolutionary biology
Background
Rate of embryonic development varies widely across species. In mammals, large-bodied species tend to have slower development rates and increased lifespan compared to small-bodied species. For example, the rate of human embryo development is about 2-3 times slower than mouse embryos even though their overall embryonic size is similar, and they undergo the same series of developmental steps. Recent work has identified the underlying cause of these differences as coming from differential rates of protein production and degradation between the two species1,2. However, what leads to these different biochemical speeds in the two species remains unknown. In this preprint, Diaz-Cuadros et al. provide the first clues towards understanding this phenomenon by investigating the role of metabolism in regulating development rates within a specific developmental process, termed the segmentation clock.
The segmentation clock is a tissue-scale rhythmic patterning system in vertebrates. Locally synchronized waves of genetic transcription sweep from the posterior of the embryo to the anterior providing spatiotemporal information to morphological somite formation. Somites are segmented tissues in the embryonic mesoderm, which give rise to the adult musculoskeletal system. Somites form in a periodic manner – about every 5 hours in human embryos and about every 2.5 hours in mouse embryos – and this periodicity emerges from periodic transcriptional waves. Given the difference in periodicity between humans and mice, the segmentation clock can be used as a powerful system to probe underlying processes driving differential rates during development. To follow periodic transcriptional waves, the authors had previously established a methodology to recapitulate the segmentation clock in vitro from pluripotent stem cells (PSC)3. Here, the same in vitro system was used to investigate the role of metabolism in the differential periodicity of the segmentation clock.
Key Findings
The authors first developed a protocol to differentiate mouse and human PSCs towards presomitic mesoderm (cell type which exhibits segmentation clock oscillations) fate under identical media conditions. An accelerated differentiation efficiency, a faster cell cycle rate and more frequent oscillations in mouse cells indicated an approximately two-fold difference in developmental rate between mouse and human cells (Fig. 1A). The observed developmental rate was cell autonomous as the segmentation clock period did not change in isolated cells nor when human cells were co-cultured at low density with mouse cells.
To investigate the source of this differential developmental rate, oxygen consumption rate (OCR) and glycolytic proton efflux rate (glycoPER), which both serve as proxies for metabolic rate, were determined. Surprisingly, both rates were similar between mouse and humans. However, as the total mass and volume of human cells were twice that of mouse cells, it was concluded that a good comparison is only possible by considering mass-specific metabolic rates, which represents the rate at which cells consume energy per gram of cell weight. Accordingly, mass-specific OCR and glycoPER were found to be twice as fast in mouse cells as in human cells (Fig. 1B). A faster mass-specific metabolic rate in mouse cells was further confirmed through multiple independent measurements of glucose consumption, lactate secretion and glutamine consumption, as well as through comparison of mouse and human neural progenitors differentiated in vitro from PSCs.
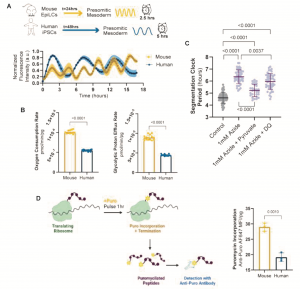
Figure 1. Key findings from the preprint. Adapted from Figs 1-4, Diaz-Cuadros et al. 2021
The authors then proceeded to test which aspect of metabolism controls developmental rate. Given that segmentation clock oscillations are known to be affected upon perturbation of respiration but not glycolysis, human presomitic mesoderm (PSM) cells were treated with small molecule inhibitors of the electron transport chain (ETC). Inhibiting ETC complexes I, III, and IV in human PSM cells resulted in a significant lengthening of the segmentation clock and damped oscillatory dynamics, while inhibition of the ATP synthase had no effects. These results suggest that ETC, rather than ATP synthase, activity is involved in the regulation of the segmentation clock. To further test the hypothesis that cellular ATP levels do not regulate the segmentation clock, the authors cultured human PSM cells in multiple conditions that increased the cellular concentration of ATP. These conditions did not shorten the segmentation clock period and increased cell cycle length, indicating slower proliferation. Therefore, the authors concluded that increased levels of ATP do not mediate the accelerated developmental rate of mouse cells.
Considering the link between a high NAD+/NADH ratio and increased proliferation rates4, the authors then tested the importance of NAD+ availability on the segmentation clock. Decreasing NAD+/NADH within human PSM cells pharmacologically or through manipulation of the cell culture medium led to a lengthening of the segmentation clock period. This effect could be rescued by regenerating NAD+ levels, suggesting the segmentation clock period depends on NAD+ levels (Fig. 1C).
Previous studies have suggested that differences in developmental rate between species was partly due to differences in protein production1,2. The authors validated this and showed that mouse PSM cells have a mass-specific translation rate that is twice as fast as human PSM cells (Fig. 1D). In addition, slowing down translation led to a significant extension of the segmentation clock period of human PSM cells. Importantly, ETC inhibition, which also lengthened the segmentation clock period, decreased translation rates, with the magnitude of the effect on clock period scaling with magnitude of translation rate inhibition. Further experiments suggested that mitochondrial activity acts upstream of translation rate to regulate the clock period. In general, a change in the clock period could also emerge from a change in protein degradation rate. However, clock oscillations were found to be much less sensitive to perturbations of proteasome activity compared to perturbations of protein production. Taken together, the authors concluded that mitochondrial activity drives the translation rate which regulates the segmentation clock period and therefore, developmental rate.
Why we like this preprint
- This is the first study that systematically compares mass-specific metabolic rates across species during embryonic development. By doing so, this work has shown Kleiber’s law, a long-standing allometric relationship between body mass and metabolic rate, to be valid during embryonic development as well.
- The discovery that developmental rate, at least in the context of segmentation clock period, is sensitive to NAD+ levels is interesting from a perspective of understanding cancer cell proliferation as well as aging-related processes, both of which are known to depend on NAD+ levels.
- This work potentially sets the stage for developing stem cell-based therapies in the future where manipulation of differentiation rate could aid in accelerating disease modeling.
Open questions for the authors
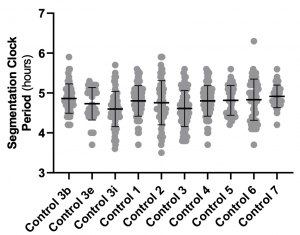
- Why are control segmentation clock periods (compare Fig. 3b, 3e and 3i) variable across experiments? If you run a statistical test across controls, would they be significantly different? On the same note, clock period under DCA-treated conditions matches the control clock period from Fig. 3b. Given this, how do you interpret changes in clock period under these conditions?
- In vivo, the electron transport chain and oxidative metabolism was previously shown to have a gradient with a higher activity in the anterior5,6. Considering results from this work, does it mean that a differential clock period between mouse and human cells can be explained as a function of metabolic rate only in the anterior?
- Do the authors plan to study whether mutations in mitochondrial/metabolic genes are linked to segmentation clock or somitogenesis defects? Have any links already been established?
- From an evolutionary perspective, it still remains unclear why different species have widely varying metabolic rates. Could the authors comment on this?
- Any speculation on why embryos from a particular species have a changed protein production rate rather than degradation rate to modulate overall developmental speed?
References:
- Matsuda, M., et al., Species-specific segmentation clock periods are due to differential biochemical reaction speeds. Science, 2020. 369(6510): p. 1450.
- Rayon, T., et al., Species-specific pace of development is associated with differences in protein stability. Science, 2020. 369(6510): p. eaba7667.
- Diaz-Cuadros, M., et al., In vitro characterization of the human segmentation clock. Nature, 2020. 580(7801): p. 113-118.
- Luengo, A., et al., Increased demand for NAD(+) relative to ATP drives aerobic glycolysis. Mol Cell, 2021. 81(4): p. 691-707.e6.
- Özbudak et al., Spatiotemporal compartmentalization of key physiological processes during muscle precursor differentiation. PNAS, 2010. 107(9): p. 4224-4229
- Oginuma et al., A gradient of glycolytic activity coordinates FGF and Wnt signaling during elongation of the body axis in amniote embryos. Dev. Cell, 2017. 40: p. 342-353
doi: https://doi.org/10.1242/prelights.30666
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the developmental biology category:
Chromatin priming and co-factor availability shape lineage response to the neuronal pioneer factor ASCL1 in pluripotency
Reinier Prosee, Jethro Lundie-Brown
Cardiac-immune microniches programme macrophage states in the regenerating heart
Theodora Stougiannou
Temporal degradation of PRC2 uncovers specific developmental dependencies
María Mariner-Faulí
Also in the evolutionary biology category:
Heatwave winners and losers: cryptic coral holobionts differ in thermal tolerance
Maitri Manjunath
No global collapse of food webs across the Permian–Triassic Mass Extinction
Theodora Stougiannou
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
preLists in the developmental biology category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
BSDB/GenSoc Spring Meeting 2024
A list of preprints highlighted at the British Society for Developmental Biology and Genetics Society joint Spring meeting 2024 at Warwick, UK.
| List by | Joyce Yu, Katherine Brown |
GfE/ DSDB meeting 2024
This preList highlights the preprints discussed at the 2024 joint German and Dutch developmental biology societies meeting that took place in March 2024 in Osnabrück, Germany.
| List by | Joyce Yu |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
2nd Conference of the Visegrád Group Society for Developmental Biology
Preprints from the 2nd Conference of the Visegrád Group Society for Developmental Biology (2-5 September, 2021, Szeged, Hungary)
| List by | Nándor Lipták |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
Single Cell Biology 2020
A list of preprints mentioned at the Wellcome Genome Campus Single Cell Biology 2020 meeting.
| List by | Alex Eve |
Society for Developmental Biology 79th Annual Meeting
Preprints at SDB 2020
| List by | Irepan Salvador-Martinez, Martin Estermann |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
Cell Polarity
Recent research from the field of cell polarity is summarized in this list of preprints. It comprises of studies focusing on various forms of cell polarity ranging from epithelial polarity, planar cell polarity to front-to-rear polarity.
| List by | Yamini Ravichandran |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EDBC Alicante 2019
Preprints presented at the European Developmental Biology Congress (EDBC) in Alicante, October 23-26 2019.
| List by | Sergio Menchero et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |
BSCB/BSDB Annual Meeting 2019
Preprints presented at the BSCB/BSDB Annual Meeting 2019
| List by | Dey Lab |
Zebrafish immunology
A compilation of cutting-edge research that uses the zebrafish as a model system to elucidate novel immunological mechanisms in health and disease.
| List by | Shikha Nayar |
Also in the evolutionary biology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
EMBO | EMBL Symposium: The organism and its environment
This preList contains preprints discussed during the 'EMBO | EMBL Symposium: The organism and its environment', organised at EMBL Heidelberg, Germany (May 2023).
| List by | Girish Kale |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
COVID-19 / SARS-CoV-2 preprints
List of important preprints dealing with the ongoing coronavirus outbreak. See http://covidpreprints.com for additional resources and timeline, and https://connect.biorxiv.org/relate/content/181 for full list of bioRxiv and medRxiv preprints on this topic
| List by | Dey Lab, Zhang-He Goh |
1
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |











 (No Ratings Yet)
(No Ratings Yet)