On-site ribosome remodeling by locally synthesized ribosomal proteins in axons
Posted on: 29 January 2019
Preprint posted on 19 December 2018
Article now published in Cell Reports at http://dx.doi.org/10.1016/j.celrep.2019.11.025
Beyond the Nucleolus: Ribosomal proteins synthesized and incorporated into ribosomes locally within axons contribute to axonal branching behavior.
Selected by Srivats VenkataramananCategories: molecular biology, neuroscience
Context:
Numerous datasets have demonstrated that the subcellular distribution of mRNA within neurons is non-uniform, and specific subsets of transcripts are enriched within the axon – distant from the main cell body and the regulatory hub of the nucleus [1]. Furthermore, recent evidence has also demonstrated that these transcripts are actually translated locally within the axon in a manner that tracks with development and the maturation of neuronal circuits [2, 3].
Among the transcripts that localize to and are translated within axons are ribosomal protein genes (RPGs) – structural components of the machinery of translation. Their presence in the axon has long been considered a conundrum, since assembly of the ribosome has canonically been thought to occur in the nucleus/nucleolus – far from the axon. The textbook view of ribosome assembly involves RPGs being translated into ribosomal proteins (RPs) in the cytoplasm, which are subsequently imported into the nucleus to assemble with the ribosomal RNA (rRNA) in the nucleolus, forming the precursors to the ribosomal subunits. Once assembled, these precursor subunits are re-exported back into the cytoplasm, where they mature and assemble upon an mRNA to form the functional ribosome [4]. Now, the magnitude of the spatial separation between RPG transcripts within the axon, and the nucleus in the cell body render this a potentially very energy-expensive process. Therefore, it has been postulated that the local translation of RPs in the axon is indicative of local function, which, given the accepted pathway of ribosome assembly described above, has been speculated to be extra-ribosomal [5].
In this preprint, the authors challenge these underlying assumptions. They demonstrate that axonally translated RPs are incorporated into ribosomes within the axon itself and that this ‘remodeling’ of the ribosome has functional consequences for axonal branching.
Key Findings:
The authors have previously developed an assay using a Boyden chamber that allows them to separate axons of retinal ganglion cells from the rest of the cell body and nucleus [3]. They find that translation of almost all of the RPGs (which were previously known to be enriched in axonal projections) transiently increases locally within the axon upon induction of axonal branching via Netrin-1. They find that increased axonal translation is dependent on a “Cis-element Upstream of the Initation Codon” (CUIC): a novel YYYYTTYC motif within the 5’ UTR of these transcripts.
Crucially, the authors demonstrate that RPs that are synthesized within the axonal projections remain in the axon. Mass-spectrometry further revealed that axonally-translated RPs –particularly RPs that are cytoplasmic facing in the ribosome – get incorporated into the axonal ribosomes. The most remarkable aspect of this result was that since the assay was performed in isolated axons that were severed from their soma, this incorporation occurred locally within the axon itself, eschewing the canonical ribosome assembly pathway which would require axonally synthesized RPs to travel to the nucleus. To my knowledge, this is the first demonstration of in situ incorporation of a RP into the ribosome without any nucleolar involvement.
Interestingly, some locally synthesized RPs like Rps4x and Rps14 are far more prone to the phenomenon than others, indicating that this isn’t full-scale ribosome biosynthesis within the axon, but rather an “add-on” to either modify or repair the axonal ribosome. This local incorporation of at least Rps4x is critical for proper functioning of the ribosome. In the absence of local Rps4x incorporation, axonal translation as measured by puromycin incorporation is severely abrogated – and this results in severe defects in axonal branching of the retinal ganglion cells.
In summary, signals that induce axonal branching during neural development induce localized gene regulatory responses within the axon. The signal induces CUIC-mediated axonal translation of RPs – some of which, such as Rps4x, get incorporated into ribosomes in their vicinity, thereby stimulating local translation of transcripts encoding for (presumably) proteins involved in axonal branching (Figure 1 – figure 7B in the manuscript).
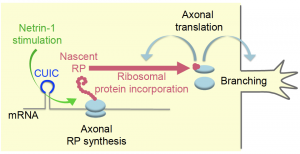
Figure 1: Axonal RP synthesis is required for local
translation of genes involved in axonal branching.
Open questions, future directions, and why I chose this preprint:
This is an extraordinary piece of work demonstrating not just axonal translation of RPs but also their local incorporation into ribosome, challenging decades of dogma. However, most of the functional assays within the manuscript have been performed using Rps4x. This makes sense, given that Rps4x shows the highest de novo incorporation into the ribosome of all of the RPs upon the induction of axonal branching (Figure 4C in the manuscript). Rps4x has previously been demonstrated to be amongst the most sub-stoichiometric of the ribosomal proteins [6]. These observations indicate at least two potential hypotheses
- Rps4x is the most intrinsically labile of the ribosomal proteins, and thus susceptible to loss from the ribosome. Ribosomes lacking Rps4x are sub-optimal in their ability to translate an mRNA, resulting in potentially stalled ribosomes and lower translation. Upon the induction of axonal branching, locally synthesized Rps4x re-associates with the ribosomes within the axon, restoring optimal ribosomal activity and the translation of transcripts involved in axonal branching and the maturation of neural circuitry.
- There are distinct populations of axonal ribosomes that either incorporate or are depleted of Rps4x. These distinct ribosomal sub populations translate different functional subsets of transcripts – and are thus required at different stages of development. The pool of transcripts that require Rps4x for their translation is involved in axonal branching, and in order to achieve their translation, Rps4x is synthesized and incorporated locally into axonal ribosomes.
Distinguishing between these hypotheses, which respectively represent repair or modification of the axonal ribosome, requires careful stoichiometric analysis of ribosomal proteins within the functional ribosome before, during and after axonal branching – an experiment that is rendered more difficult by limitations of the starting material, as acknowledged by the authors. However, this system provides a potentially tractable model to address a critical question in the burgeoning study of heterogeneous ribosomes: are ribosomes truly ‘specialized’ to translate subsets of transcripts [7], or are the observed gene expression effects of specific RP deletions simply the result of a hypomorphic ribosome unsuccessfully navigating a (tissue-specific) hierarchal transcript landscape with different intrinsic initation and elongation rates [8]?
Two questions remain on the RNA front. Firstly, which transcripts are locally translated by the Rps4x containing axonal ribosomes that contribute to branching? Do they also contain the CUIC sequence motif enriched in RPG transcripts? Are they among the transcripts involved in neurogenesis/neuron projection differentiation/morphogenesis that harbor the CUIC motif (Figure 2C)? If so, that speaks to an interesting positive feedback loop wherein the presence of Rps4x containing ribosomes results in the translation of more Rps4x. How might the loop be terminated upon the cessation of axonal branching? Secondly, what is the trans-factor that enhances the local translation of CUIC containing mRNAs upon the induction of branching?
Finally, taking a birds-eye view, how generalizable is this phenomenon? Non-uniform transcript distribution within the cytoplasm is certainly not unique to neurons. Nor, in fact, is localized translation within cellular domains. One can imagine virtually any polarized cell (such as in the gut epithelium) leveraging similar mechanisms to achieve both energy economy and rapid responses [9]. Long gone are the days of envisioning the cytoplasm as a pool under the tyranny of Brownian diffusion. It is in fact a thriving city, a loose amalgam of neighborhoods, each with their own rules and personalities, and each worthy of intrepid exploration.
References:
- Taliaferro, J.M., et al., Distal Alternative Last Exons Localize mRNAs to Neural Projections. Mol Cell, 2016. 61(6): p. 821-33.
- Shigeoka, T., et al., Dynamic Axonal Translation in Developing and Mature Visual Circuits, in Cell. 2016. p. 181-92.
- Cagnetta, R., et al., Rapid Cue-Specific Remodeling of the Nascent Axonal Proteome. Neuron, 2018. 99(1): p. 29-46.e4.
- Lafontaine, D.L., Noncoding RNAs in eukaryotic ribosome biogenesis and function. Nat Struct Mol Biol, 2015. 22(1): p. 11-9.
- Warner, J.R. and K.B. McIntosh, How common are extraribosomal functions of ribosomal proteins? Mol Cell, 2009. 34(1): p. 3-11.
- Slavov, N., et al., Differential Stoichiometry among Core Ribosomal Proteins, in Cell Rep. 2015. p. 865-73.
- Genuth, N.R. and M. Barna, Heterogeneity and specialized functions of translation machinery: from genes to organisms. Nat Rev Genet, 2018. 19(7): p. 431-452.
- Mills, E.W. and R. Green, Ribosomopathies: There’s strength in numbers. Science, 2017. 358(6363).
- Moor, A.E., et al., Global mRNA polarization regulates translation efficiency in the intestinal epithelium. Science, 2017. 357(6357): p. 1299-1303.
doi: https://doi.org/10.1242/prelights.7988
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the molecular biology category:
Chromatin priming and co-factor availability shape lineage response to the neuronal pioneer factor ASCL1 in pluripotency
Reinier Prosee, Jethro Lundie-Brown
Temporal degradation of PRC2 uncovers specific developmental dependencies
María Mariner-Faulí
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
Also in the neuroscience category:
Generalization and extinction of learned fear alter primary sensory input to the brain
Kathleen Ngo et al.
Fear conditioning biases olfactory sensory neuron frequencies across generations
Yuwei Wu et al.
Chromatin priming and co-factor availability shape lineage response to the neuronal pioneer factor ASCL1 in pluripotency
Reinier Prosee, Jethro Lundie-Brown
preLists in the molecular biology category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |
Also in the neuroscience category:
preLighters’ choice – Handpicked DevBio preprints
preLighters with expertise across developmental and stem cell biology have nominated a few developmental biology (and related) preprints they’re excited about and explain in a few paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Theodora Stougiannou et al. |
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |











 (No Ratings Yet)
(No Ratings Yet)