Patient-derived ovarian cancer organoids mimic clinical response and exhibit heterogeneous inter- and intrapatient drug responses
Posted on: 31 December 2019 , updated on: 27 January 2020
Preprint posted on 15 December 2019
Show yourself: patient-derived organoids meet epithelial ovarian cancer (OC)—using organoids to enable personalised therapy
Selected by Zhang-He GohCategories: cancer biology, pharmacology and toxicology, systems biology
Background of preprint: secrets deep inside
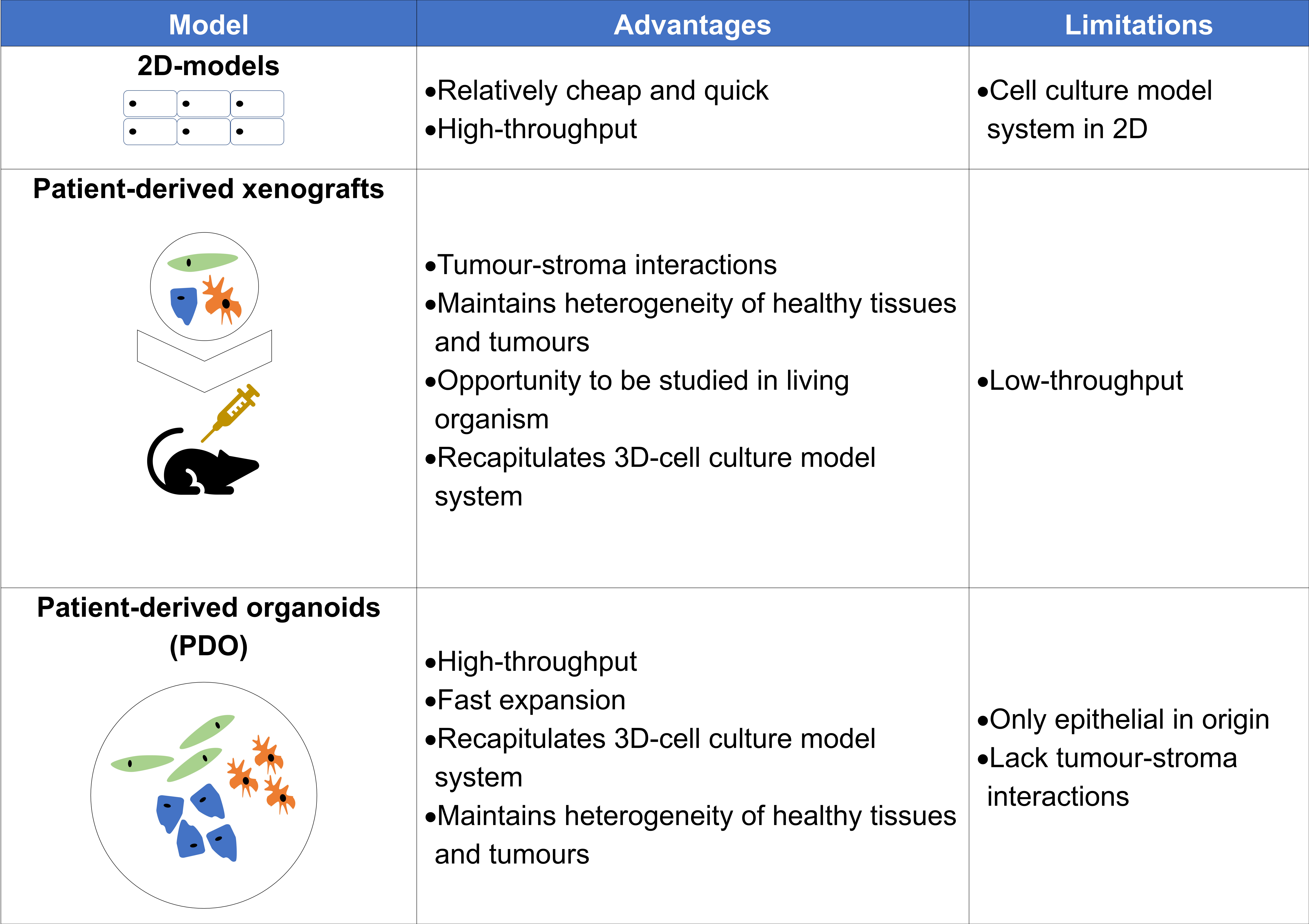
Two major problems are currently associated with epithelial ovarian cancer (OC): chemotherapy resistance and poor survival. Despite the development of various targeted therapies and surgical techniques for tumour resection, overall survival remains poor. OC has traditionally been studied using 2D-cell lines and xenografts, each with their respective advantages and limitations (Table 1). The need for alternative models has led to the development of patient-derived organoids (PDOs). Touted by de Witte et al. as “3D-cell culture (models) that (maintain) the cellular heterogeneity of healthy tissues and tumours”, PDOs have been described to allow for both high-throughput drug screens [1] while preserving the genomic features of the tumours from which they were derived [2-4]. By accounting for genetic heterogeneity which often frustrates efforts in overcoming resistance to anticancer agents [5], PDOs may enable researchers and clinicians better establish correlations between drug responses observed in PDOs and in humans.
While some work has been performed on establishing this correlation [3,4], the evidence has been relatively anecdotal. In their preprint, de Witte et al. systematically assess the correlation between OC PDOs and patients’ clinical response to chemotherapeutics.
Table 1. Summary of models’ advantages and limitations.
Key findings of preprint: PDOs have genomic “memory”
First, de Witte et al. characterised 36 PDOs, derived from 23 patients with different histological subtypes of OC. By comparing the genomic profiles of these PDOs and their tumours of origin, the authors found that the PDOs are enriched in tumour cells; in contrast, tumour samples are heterogeneous in containing a mix of tumour and normal cells.
The preprint authors then correlated the drug responses between PDOs and patients’ clinical response. de Witte et al. exposed 7 PDOs (derived from 5 patients) to carboplatin and paclitaxel combination treatment in vitro (preprint Figure 1). By expanding this to other PDOs to make up a total of 36 PDOs, de Witte et al. found that PDOs exhibit inter-patient drug response heterogeneity, which correlated partially with their genetic makeup. The preprint authors made two observations:
- Responses to carboplatin, paclitaxel, and gemcitabine were divergent.
- Responses correlated with OC histological subtype.
Finally, de Witte et al. found that in addition to exhibiting inter-patient drug response heterogeneity, PDOs also exhibited intra-patient drug response heterogeneity. Specifically, among the 21 PDOs derived from 8 patients*, de Witte et al. observed low drug response variability across biological replicates, but all related PDOs exhibited differential responses to at least one drug (preprint Figure 4). Among related PDOs, the authors also characterised genome-wide heterogeneity using structural variants (SVs)—mutations that affect large segments of DNA larger than 100 bp [6], copy number alterations (CNAs), and single nucleotide variants (SNVs). Interestingly, none of these were associated with phenotypic heterogeneity, i.e. variation in drug response, on a genome-wide analysis. In contrast, CNVs and SNVs were observed to correlate with phenotypic heterogeneity on an individual gene level.**
What I like about this preprint: the answer we’ve waited for?
Last year, I wrote about the development of bile duct and liver organoid models that would accelerate research on biliary tract and liver toxicities. In line with this theme of organoid development, I selected this preprint in my first preLight this year to highlight the potential applications of organoids in the clinic.
By investigating the use of PDOs as a diagnostic tool in an era of personalised medicine in their preprint, de Witte et al. help to extend the capabilities of organoid technology. Specifically, the authors describe how OC PDOs meet the three criteria of model systems in the clinic:
- PDOs genetically resemble the original tumour from which they are derived;
- PDO drug responses generally reflect patients’ clinical response;
- PDO establishment and drug screening can be performed rapidly.
Therefore, the preprint authors demonstrate the importance of conducting histopathological assessments. These could be used to support current gold-standard measures of patient response commonly used in clinical trials, such as CA-125 and RECIST criteria. I also look forward to the preprint authors’ plan for a prospective trial involving both drug tests on PDOs with simultaneous monitoring of clinical response using the same drug combinations.
Future directions: throw yourself into something new
Given the highly nascent nature of this work, some caution may be advised as we venture into the unknown: that de Witte et al. only found a partial link between genomic and phenotypic heterogeneity is particularly telling. The authors point out that until a robust correlation has been established between genetic and functional testing, clinical decisions may have to be based on both genetic and functional tests, rather than just the use of genetic tests that is so commonplace today. Indeed, in their preprint, de Witte et al. also call for follow-up studies with increased sample sizes and deeper sequencing to detect associations between genetic and functional heterogeneity.
Other questions will also need to be answered in bringing these PDO models from the bench to the bedside. Further upstream, pharmacokinetic (PK)-pharmacodynamic (PD) correlations using both in vitro and in vivo models will need to be established for in vitro findings to translate into decisions on drug selection and dosing. Clinically, cancer regimens are often complicated and can involve cocktails of multiple drugs with different dosing schedules; while the use of these PDOs may enhance the efficacy of existing medicines, their effective use may add another layer of complexity in patients’ cancer treatments. Whether this will affect patient compliance and increase the risk of medication-related errors also remains to be seen.
Then there are cost considerations. Some things never change: as government institutions battle rising healthcare costs [7,8], concerns about financial toxicity continue to impede the full implementation of personalised medicine in the clinic. Whether governmental healthcare agencies will deem such undertakings to be value for money is yet another question.
Questions for authors
- In the Materials and Methods section on in vitro PDO drug response testing and data-analysis, you wrote that “PDOs were exposed to drugs in varying concentrations and to controls” (page 4) [4]. Could you briefly elaborate on how the in vitro concentrations of drugs were selected? For instance, in the Results section (preprint page 8), “seven PDOs… were exposed to carboplatin and paclitaxel combination treatment in vitro”; how were the respective concentrations of these assays decided?
References
[1] Bleijs M, van de Wetering M, Clevers H, Drost J, Xenograft and organoid model systems in cancer research, The EMBO Journal 38(15) (2019) e101654.
[2] Jabs J, Zickgraf FM, Park J, Wagner S, Jiang X, Jechow K, Kleinheinz K, Toprak UH, Schneider MA, Meister M, Spaich S, Sütterlin M, Schlesner M, Trumpp A, Sprick M, Eils R, Conrad C, Screening drug effects in patient-derived cancer cells links organoid responses to genome alterations, Mol Syst Biol 13(11) (2017) 955-955.
[3] Hill SJ, Decker B, Roberts EA, Horowitz NS, Muto MG, Worley MJ, Jr., Feltmate CM, Nucci MR, Swisher EM, Nguyen H, Yang C, Morizane R, Kochupurakkal BS, Do KT, Konstantinopoulos PA, Liu JF, Bonventre JV, Matulonis UA, Shapiro GI, Berkowitz RS, Crum CP, D’Andrea AD, Prediction of DNA Repair Inhibitor Response in Short-Term Patient-Derived Ovarian Cancer Organoids, Cancer Discov 8(11) (2018) 1404-1421.
[4] Kopper O, de Witte CJ, Lõhmussaar K, Valle-Inclan JE, Hami N, Kester L, Balgobind AV, Korving J, Proost N, Begthel H, van Wijk LM, Revilla SA, Theeuwsen R, van de Ven M, van Roosmalen MJ, Ponsioen B, Ho VWH, Neel BG, Bosse T, Gaarenstroom KN, Vrieling H, Vreeswijk MPG, van Diest PJ, Witteveen PO, Jonges T, Bos JL, van Oudenaarden A, Zweemer RP, Snippert HJG, Kloosterman WP, Clevers H, An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity, Nature Medicine 25(5) (2019) 838-849.
[5] Vasan N, Baselga J, Hyman DM, A view on drug resistance in cancer, Nature 575(7782) (2019) 299-309.
[6] Lee AY, Ewing AD, Ellrott K, Hu Y, Houlahan KE, Bare JC, Espiritu SMG, Huang V, Dang K, Chong Z, Caloian C, Yamaguchi TN, Barnes BD, Birol I, Chen X, Chiu R, Cox AJ, Ding L, Fritz MHY, Grigoriev A, Hach F, Kawash JK, Korbel JO, Kruglyak S, Liao Y, McPherson A, Nip KM, Rausch T, Sahinalp SC, Sarrafi I, Saunders CT, Schulz-Trieglaff O, Shaw R, Shi W, Smith SD, Song L, Wang D, Ye K, Kellen MR, Chen K, Norman TC, Friend SH, Guinney J, Stolovitzky G, Haussler D, Margolin AA, Stuart JM, Boutros PC, Participants I-TDSMCC, Combining accurate tumor genome simulation with crowdsourcing to benchmark somatic structural variant detection, Genome Biology 19(1) (2018) 188.
[7] Teerawattananon Y, Teo YY, Dabak S, Rattanavipapong W, Isaranuwatchai W, Wee H-L, Luo N, Morton A, Tackling the 3 Big Challenges Confronting Health Technology Assessment Development in Asia: A Commentary, Value in Health Regional Issues 21 (2020) 66-68.
[8] Dombrádi V, Pitini E, van El CG, Jani A, Cornel M, Villari P, Gray M, Bíró K, Value-based genomic screening: exploring genomic screening for chronic diseases using triple value principles, BMC Health Serv Res 19(1) (2019) 823-823.
Corrigendum:
*This section originally referred to 40 PDOs derived from 8 patients, when in fact this number should be 21 PDOs derived from 8 patients.
**This section now clarifies that, on an individual gene level, CNVs and SNVs were associated with phenotypic heterogeneity; in contrast, the genome-wide total number of SVs, CNVs, and SNVs were not associated with phenotypic heterogeneity.
doi: https://doi.org/10.1242/prelights.16031
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the cancer biology category:
A Novel Chimeric Antigen Receptor (CAR) - Strategy to Target EGFRVIII-Mutated Glioblastoma Cells via Macrophages
Dina Kabbara
Taxane-Induced Conformational Changes in the Microtubule Lattice Activate GEF-H1-Dependent RhoA Signaling
Vibha SINGH
ROCK2 inhibition has a dual role in reducing ECM remodelling and cell growth, while impairing migration and invasion
Sharvari Pitke
Also in the pharmacology and toxicology category:
Small Molecule Agonists of TREM2 Reprogram Microglia and Protect Synapses in Human Alzheimer’s Models
Dina Kabbara
Snake venom metalloproteinases are predominantly responsible for the cytotoxic effects of certain African viper venoms
Daniel Osorno Valencia
Cryo-EM reveals multiple mechanisms of ribosome inhibition by doxycycline
Leonie Brüne
Also in the systems biology category:
Human single-cell atlas analysis reveals heterogeneous endothelial signaling
Charis Qi
Longitudinal single cell RNA-sequencing reveals evolution of micro- and macro-states in chronic myeloid leukemia
Charis Qi
Environmental and Maternal Imprints on Infant Gut Metabolic Programming
Siddharth Singh
preLists in the cancer biology category:
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
March in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) cancer biology 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics and genomics 6) other
| List by | Girish Kale et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
Single Cell Biology 2020
A list of preprints mentioned at the Wellcome Genome Campus Single Cell Biology 2020 meeting.
| List by | Alex Eve |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Anticancer agents: Discovery and clinical use
Preprints that describe the discovery of anticancer agents and their clinical use. Includes both small molecules and macromolecules like biologics.
| List by | Zhang-He Goh |
Biophysical Society Annual Meeting 2019
Few of the preprints that were discussed in the recent BPS annual meeting at Baltimore, USA
| List by | Joseph Jose Thottacherry |
Also in the pharmacology and toxicology category:
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
COVID-19 / SARS-CoV-2 preprints
List of important preprints dealing with the ongoing coronavirus outbreak. See http://covidpreprints.com for additional resources and timeline, and https://connect.biorxiv.org/relate/content/181 for full list of bioRxiv and medRxiv preprints on this topic
| List by | Dey Lab, Zhang-He Goh |
1
Drug use in special populations
Any drugs that are being used in special populations: Patients with liver and kidney failure, in paediatrics, in geriatrics, and in pregnant or lactating patients. Includes the discovery of factors that could potentially affect drug use in these special populations.
| List by | Zhang-He Goh |
Toxicology of toxicants, existing therapeutics, and investigational drugs
Preprints that describe the toxicology of environmental pollutants and existing and upcoming drugs. Includes both toxicokinetics and toxicodynamics, as well as technological improvements that will help in the characterisation of this field.
| List by | Zhang-He Goh |
Antimicrobials: Discovery, clinical use, and development of resistance
Preprints that describe the discovery of new antimicrobials and any improvements made regarding their clinical use. Includes preprints that detail the factors affecting antimicrobial selection and the development of antimicrobial resistance.
| List by | Zhang-He Goh |
Anticancer agents: Discovery and clinical use
Preprints that describe the discovery of anticancer agents and their clinical use. Includes both small molecules and macromolecules like biologics.
| List by | Zhang-He Goh |
Advances in Drug Delivery
Advances in formulation technology or targeted delivery methods that describe or develop the distribution of small molecules or large macromolecules to specific parts of the body.
| List by | Zhang-He Goh |
Also in the systems biology category:
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
Single Cell Biology 2020
A list of preprints mentioned at the Wellcome Genome Campus Single Cell Biology 2020 meeting.
| List by | Alex Eve |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |











 (No Ratings Yet)
(No Ratings Yet)