Deterministic and probabilistic fate decisions co-exist in a single retinal lineage
Posted on: 15 November 2022 , updated on: 22 August 2023
Preprint posted on 11 August 2022
Article now published in The EMBO Journal at http://dx.doi.org/10.15252/embj.2022112657
To be or not to be a photoreceptor: cell fate choices in the developing zebrafish retina are deterministic and probabilistic.
Selected by Laura CelottoCategories: developmental biology, neuroscience
Updated 22 August 2023 with a postLight by Laura Celotto
“The EMBO Journal” has now published the insightful work below, which even made the cover of the journal last 17 July 2023. On the cover, you can appreciate the beauty of the zebrafish retina, which looks like a carpet of well-ordered layers, each of them hosting specific subsets of cells.
In the published paper, the authors have added two experiments that expand the findings of the preprint, without changing its main conclusions.
The first experiment attempts to answer one of the questions I also raise in my highlight on the preprint: How do you explain the thickness reduction of the ganglion cell layer upon Prdm1a knock down? As a brief recap, Prdmn1a is a transcription factor that determines photoreceptor fate in the developing zebrafish retina. In the preprint, the authors could show that injection of Prdmn1a morpholinos, which knock down the protein, decreased the thickness of not only the photoreceptor layer, but also of the ganglion cell layer.
In a new experiment of the published paper, the authors looked for signs of apoptosis in controls as well as Prdmn1a-injected zebrafish to check whether the injection of Prdmn1a morpholinos leads to unspecific cell death. Apoptosis is a form of programmed cell death, characterized by the activation of caspases, which are proteases – that is, proteins that “eat” other proteins. Among caspases, activated caspase-3 is considered the hallmark of apoptotic cells. The authors performed antibody staining against activated-caspase 3 in control as well as Prdmn1a knock down retinae. They observed few activated-caspase 3-positive cells in all layers of controls and Prdmn1a-knock down retinae, with no significant difference in the number of apoptotic cells between the two groups. Hence, they could conclude that apoptosis does not contribute to the thickness reduction of the photoreceptor and ganglion cell layers upon Prdmn1a knock down. Then, the authors speculated that Prdmn1a knock down could induce a decrease in the overall number of Atoh7-positive progenitors and derived neurons, with the subsequent shrinkage of the ganglion cell layer as a result. However, the authors had already shown in the preprint that the Atoh7-progenitors generate a deterministic branch (producing photoreceptors) and a probabilistic branch that produces retinal ganglion cells as well as horizontal and amacrine cells. If it were true that Prdmn1a knock down induces the decrease of Atoh7-positive progenitors, why would such decrease affect only the ganglion cell layer and not the inner nuclear layer, where horizontal and amacrine cells reside? The authors did not observe any shrinkage of the inner nuclear layer upon Prdmn1a knock down in the published paper.
The second, novel experiment in the published paper is a characterization of the outer nuclear layer upon knock down of both Atoh7 and Ptf1a transcription factors, as compared to wild type controls. The authors observed an increased number of densely packed photoreceptor precursors in the outer nuclear layer of double morphants as compared to control animals. Moreover, photoreceptors in double morphants had a more elongated morphology than those in controls, but the expression of the photoreceptor differentiation marker Zpr1 did not change between Atoh7- and Ptf1a- double knock down and control retinae. However, Zpr1 is a marker for only a subset of cone photoreceptors. I am therefore wondering whether testing for additional markers of photoreceptor differentiation (e.g. the pan-cone marker Gnat2 as well as markers of rod differentiation) would unravel more subtle changes of photoreceptor production in the double morphant retinae, as compared to controls. On the other hand, Gnat2 and rod differentiation markers usually come up much later in retinal development, and might not be anyway detected at the time points studied by the authors in the current paper. Hence, further experiments are necessary to answer my questions, which leads me to highlight here that scientific research is a never-ending quest for answers. Sometimes you are lucky, and you think that you have answered your original question. However, when you look closer, further questions always arise and await testing by further scientists that will open new doors, which will be crossed, further opened, or slammed shut by other scientists.
And this is both the beauty and the curse of research: small steps that open new doors that open new doors, in the endless attempt to come a little closer to…the truth of Nature.
Will we ever be able to reach such truth? We really do not know, but- in the meantime- I encourage all of you to enjoy the journey.
WHY I CHOSE THIS PREPRINT
Many studies in developmental biology have investigated the development of the vertebrate retina in zebrafish, thanks to the transparency of the larvae, which are suitable to live imaging microscopy. However, this preprint by Nerli and colleagues represents the first study that combines in vivo imaging of the developing retina with in silico simulations to explain cell fate choices of retinal progenitor cells (RPCs) that express the transcription factor Atoh7. I selected this preprint because I enjoyed the accurate characterization of fate commitment at a single cell level, a novelty in the study of retinogenesis, since previous works have dissected fate choices only at RPC population or clonal level. Moreover, there is a plot-twist in the story: Atoh7-RPCs have the potential to generate more neuronal types than previously thought… Do you want to know more? Keep reading then…
KEY FINDINGS
Atoh7-RPCs generate a photoreceptor and a second cell with variable identity across time
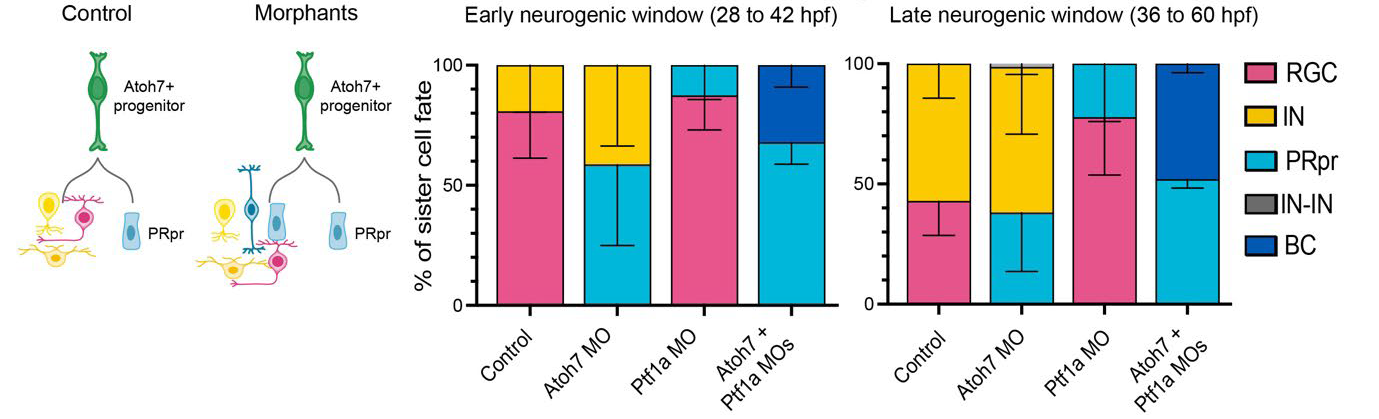
The authors used a transgenic fish line expressing a fluorescent reporter under the control of atoh7, one of the earliest neurogenic markers during retinogenesis. In this way, they could track cell divisions of single Atoh7-positive RPCs (Atoh7-RPCs) using live imaging of zebrafish embryos between 28 hours post-fertilization (hpf, the beginning of neurogenesis), and 60 hpf. They found that Atoh7-RPCs divided and generated two daughter cells with different fates. One daughter cell was always a precursor of photoreceptors, the light-sensing cells of the retina. Photoreceptor precursors further divided symmetrically to produce two photoreceptors. The second daughter of the Atoh7-RPCs, instead, could be a retinal ganglion cell, which is the output neuron of the retina, or a horizontal or an amacrine cell, the main inhibitory neurons in the retina. The probability that the Atoh7-RPCs would produce a retinal ganglion cell rather than an inhibitory neuron changed across time in neurogenesis. Indeed, RPCs during early (28-42 hpf) neurogenesis generated a photoreceptor precursor and most likely a retinal ganglion cell. Conversely, RPCs during late (36-60 hpf) neurogenesis generated a photoreceptor precursor and most likely an inhibitory neuron. Hence, Atoh7 RPCs differentiate along two branches: a deterministic branch that always produces photoreceptors, and a probabilistic branch that generates different cell types (a retinal ganglion cell or an inhibitory neuron) with different probabilities across time.
Faithful photoreceptor genesis is necessary for correct lamination of the retina
Next, the authors perturbed the outcome of Atoh7-RPC divisions by using morpholinos (short, antisense oligonucleotides) to knock down the expression of distinct fate determinants of the deterministic and probabilistic branches. Knocking down of Prdm1a, which is a transcription factor necessary for photoreceptor determination, impaired the deterministic branch of the Atoh7 lineage, so that photoreceptors were never produced at any time in retinogenesis. Indeed, interfering with the deterministic branch of the Atoh7-RPC division resulted in the production of a cell that initially looked like a photoreceptor precursor, but never divided further, nor differentiated to photoreceptors. Moreover, severe impairment of retina lamination occurred, with reduction of the photoreceptor layer and of the retinal ganglion cell layer. Conversely, perturbation of the probabilistic branch resulted in changed probabilities of the retinal neurons produced by Atoh7-RPC divisions, without affecting retinal lamination. Indeed, knock down of Atoh7 depleted retinal ganglion cells, but increased genesis of inhibitory neurons as well as of photoreceptors, with no significant effect on the overall thickness of the retina. Moreover, knock down of Ptf1a, a transcription factor necessary for the development of inhibitory neurons, resulted in decreased genesis of horizontal and amacrine cells, but increased production of retinal ganglion cells as well as of photoreceptors (Figure 1).
So, what do you think would happen if both Atoh7 and Ptf1a were knocked down?
Plot twist: Atoh7 RPCs have the potential to generate all retinal neurons, even bipolar cells!
Previous work1 and the current preprint show that bipolar cells, the last-born neurons in the retina, usually come from RPCs that never express Atoh7. Indeed, right before neurogenesis, a RPC divides to generate an Atoh7-positive lineage, including the probabilistic and deterministic branches, and an Atoh7-negative lineage that generates precursors of bipolar cells expressing the transcription factor Vsx1. However, the present study shows that knock down of Atoh7 and Ptf1a proteins in Atoh7-RPCs produced photoreceptor precursors as well as bipolar cells throughout the neurogenic window (Figure 1). If Atoh7-RPCs were not multipotent, they would never produce any bipolar cells upon Atoh7 and Ptf1a depletion. Hence, the authors concluded that Atoh7-RPCs potentially can generate all retinal neurons, including bipolar cells, but are eventually competent to produce only retinal ganglion, horizontal, amacrine and photoreceptor cells.
Figure 1. Interference with the probabilistic branch of Atoh7-RPCs. In control retinae, Atoh7-RPCs always generated a photoreceptor precursor (PRpr, deterministic branch) and another cell that could be either a retinal ganglion cell or an inhibitory neuron (IN) (probabilistic branch). In the morphant retina, where Atoh7 had been knocked down (Atoh7 MO), PRpr and IN production increased as compared to control during early and late neurogenesis. In Ptf1a MO retinae, RGC and PRpr genesis increased as compared to control during early and late neurogenesis. In the Atoh7+Ptf1a double MO retinae, PRpr genesis increased and BC production appeared. Figure modified from the original preprint.
A simple network of factors governs RPC fate choices
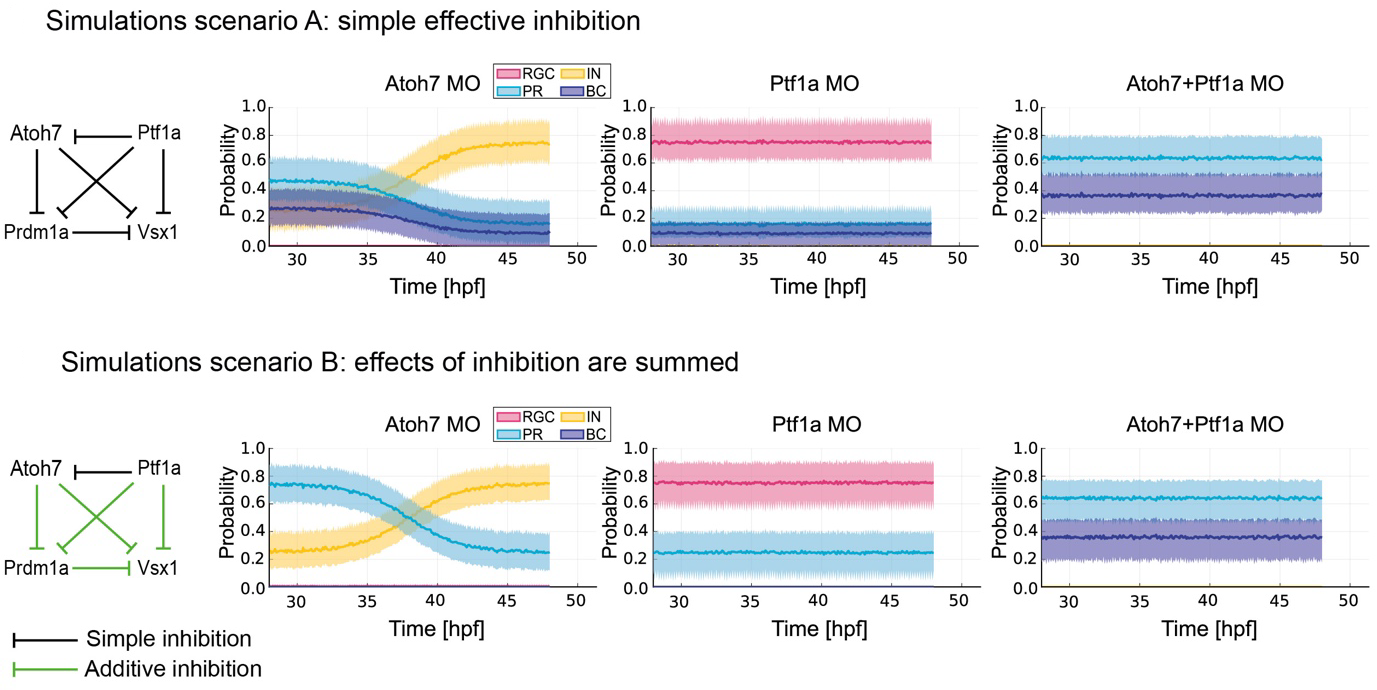
Finally, the authors developed a computational model to investigate what type of gene regulatory network involving Atoh7, Ptf1a, Prdm1a and Vsx1 would be sufficient to predict cell fate choices of Atoh7-RPCs (Figure 2). They tested two possible in silico scenarios. In the first one, it was assumed that a single transcription factor was enough to effectively inhibit the expression of a downstream factor, and, consequently, the related fate acquisition. The fate outcomes predicted by this first model, however, did not match the observed outcomes of cell divisions: indeed, the first simulation predicted the genesis of bipolar cells even in the single knock down for Atoh7 or Ptf1a, which was never detected by live imaging experiments. In other words, if it were true that, for instance, Atoh7 alone would effectively inhibit Vsx1, then we would expect bipolar cell production upon Atoh7-single knock down (Figure 2, scenario A). This simulation did not match the live imaged fate outcomes observed in the retinae of the single knock downs, nor the neuronal proportions observed in wild type embryos. In the second scenario, it was assumed that a combination of transcription factors have to exert a summed inhibition on a downstream factor to prevent the acquisition of a certain fate (Figure 2, scenario B). For instance, Atoh7 and Ptf1a were predicted to sum their inhibitions against the downstream factor Vsx1, preventing the bipolar cell fate acquisition by the Atoh7-RPCs. Indeed, it was necessary to experimentally knock down both Atoh7 and Ptf1a to observe bipolar cell genesis, while neither the Atoh7-single knock down, nor the Ptf1a-single knock down were enough to generate bipolar cells in the Atoh7-RPC, probabilistic branch (compare Figure 1 ad Figure 2, scenario B).
Figure 2. A summed inhibition is required to prevent the acquisition of a certain cell fate. In the first simulations scenario (scenario A), each transcription factor alone is able to effectively inhibit a downstream transcription factor and the acquisition of the cell fate specified by the inhibited factor. This model predicts bipolar cell (BC) production in the Atoh7-single morphant (MO) retinae, an outcome never observed with live imaging experiments (see Figure 1). In the second simulations scenario (scenario B), the effects of inhibition from more than one transcription factor are summed. This model predicts increased probability of photoreceptor (PR) and inhibitory neurons (IN) production in the Atoh7-single MO retinae, of photoreceptor and retinal ganglion cell (RGC) production in the Ptf1a-single MO retinae and of photoreceptor and bipolar cell production in the Atoh7+Ptf1a-double MO retinae. The predictions made by the simulation scenario B match the observed outcomes of live imaging experiments. Figure modified from the original preprint.
QUESTIONS FOR THE AUTHORS
1. You use morpholinos to prevent the expression of transcription factors during retinogenesis, but how specific is the action of morpholinos? Did you consider the potential off-target effects of these morpholinos? Did you think of using a conditional knock out approach to answer your questions?
2. Do you have a hypothesis about the molecular signal that starts the Atoh7 lineage? That is, did you obtain any indication about what molecular cue triggers the “firing” of Atoh7 in RPCs?
3. Did you further characterize the “aberrant” photoreceptor precursor upon Prdm1a knock down, for instance using immunohistochemistry or other approaches?
4. How do you explain the thickness reduction of the ganglion cell layer upon Prdm1a knock down?
REFERENCES
1. Vitorino, M., Jusuf, P.R., Maurus, D., Kimura, Y., Higashijima, S., and Harris, W.A. (2009). Vsx2 in the zebrafish retina: restricted lineages through derepression. Neural Develop. 4, 14. 10.1186/1749-8104-4-14.
doi: https://doi.org/10.1242/prelights.33096
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the developmental biology category:
Cell-intrinsic compliance mechanism enables release of tensile stress to prevent tissue rupture
Ruoheng Li
Cooperation between cortical and cytoplasmic forces shapes planar 4-cell stage embryos
Corentin Mollier, Shivani Dharmadhikari
A drought stress-induced MYB transcription factor regulates pavement cell shape in leaves of European aspen (Populus tremula)
Jeny Jose
Also in the neuroscience category:
Consciously detecting and recognizing a past visual word after its sensory trace is gone
Maitri Manjunath
Exogenous Amyloid Sequences: Their Role in Amyloid-Beta Heterotypic Aggregation
Cyntia Alves Conceição, Marcus Oliveira
Human Assembloid Model of Emergent Neurotropic Enteroviruses
Mitchell Sarmie
preLists in the developmental biology category:
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
BSDB/GenSoc Spring Meeting 2024
A list of preprints highlighted at the British Society for Developmental Biology and Genetics Society joint Spring meeting 2024 at Warwick, UK.
| List by | Joyce Yu, Katherine Brown |
GfE/ DSDB meeting 2024
This preList highlights the preprints discussed at the 2024 joint German and Dutch developmental biology societies meeting that took place in March 2024 in Osnabrück, Germany.
| List by | Joyce Yu |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
2nd Conference of the Visegrád Group Society for Developmental Biology
Preprints from the 2nd Conference of the Visegrád Group Society for Developmental Biology (2-5 September, 2021, Szeged, Hungary)
| List by | Nándor Lipták |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
EMBL Conference: From functional genomics to systems biology
Preprints presented at the virtual EMBL conference "from functional genomics and systems biology", 16-19 November 2020
| List by | Jesus Victorino |
Single Cell Biology 2020
A list of preprints mentioned at the Wellcome Genome Campus Single Cell Biology 2020 meeting.
| List by | Alex Eve |
Society for Developmental Biology 79th Annual Meeting
Preprints at SDB 2020
| List by | Irepan Salvador-Martinez, Martin Estermann |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
Cell Polarity
Recent research from the field of cell polarity is summarized in this list of preprints. It comprises of studies focusing on various forms of cell polarity ranging from epithelial polarity, planar cell polarity to front-to-rear polarity.
| List by | Yamini Ravichandran |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EDBC Alicante 2019
Preprints presented at the European Developmental Biology Congress (EDBC) in Alicante, October 23-26 2019.
| List by | Sergio Menchero et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |
Pattern formation during development
The aim of this preList is to integrate results about the mechanisms that govern patterning during development, from genes implicated in the processes to theoritical models of pattern formation in nature.
| List by | Alexa Sadier |
BSCB/BSDB Annual Meeting 2019
Preprints presented at the BSCB/BSDB Annual Meeting 2019
| List by | Dey Lab |
Zebrafish immunology
A compilation of cutting-edge research that uses the zebrafish as a model system to elucidate novel immunological mechanisms in health and disease.
| List by | Shikha Nayar |
Also in the neuroscience category:
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
SDB 78th Annual Meeting 2019
A curation of the preprints presented at the SDB meeting in Boston, July 26-30 2019. The preList will be updated throughout the duration of the meeting.
| List by | Alex Eve |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Young Embryologist Network Conference 2019
Preprints presented at the Young Embryologist Network 2019 conference, 13 May, The Francis Crick Institute, London
| List by | Alex Eve |











 (No Ratings Yet)
(No Ratings Yet)