Optical determination of absolute membrane potential
Posted on: 15 March 2019 , updated on: 20 March 2019
Preprint posted on 14 January 2019
New fluorescence lifetime-based approach optically determines absolute membrane potential with 20-fold greater accuracy than ever before.
Selected by James MarchantCategories: bioengineering, molecular biology, physiology
Background
Membrane potential (Vmem) is an essential component of cellular physiology and is an essential signaling cue of cellular processes such as migration and cellular division. Vmem states in cell populations are largely uncharacterized due to the difficulty of patch clamp at specific stages but reports suggest that Vmemplays a role in phases of the cell cycle1,2. Although patch clamp electrophysiology is traditionally used for measuring Vmem,this method is highly invasive and time consuming. The possibility of modifying the signals under study with introduction of pipette solution into the cell and the low throughput of patch clamping created the need for alternative methods for determining absolute membrane potential3,4.
Key findings
Lazzari-Deanet al have developed an interesting new method for optically quantifying absolute membrane potential in living cells using a fluorescent lifetime-based approach with single cell resolution. Lifetime-voltage relationships were established using VoltageFluor-fluorescent lifetime images (VF-FLIM) and simultaneous electrophysiology reporting a linear relationship for absolute Vmemand a 20-fold improved accuracy over previous optical techniques. Single cell or cell populations can be measured, increasing spatial resolution 100-fold compared to patch clamping.
Evaluation of VF-FILM across cell lines
VF-FILM has high sensitivity (3.1 – 3.7ps/mV) and short average 0mV lifetime (1.78 – 1.87 ns). VF-FILM can be used across an array of cell types commonly used in electrophysiological studies (Fig 1) and can reliably be used in cell populations requiring only single-point calibration. All cells showed linear relationship between VF fluorescent lifetime and Vmem with a voltage resolution of 5mV or better. Not only does this mean a higher throughput, but also an increased spatial resolution compared to patch clamp.
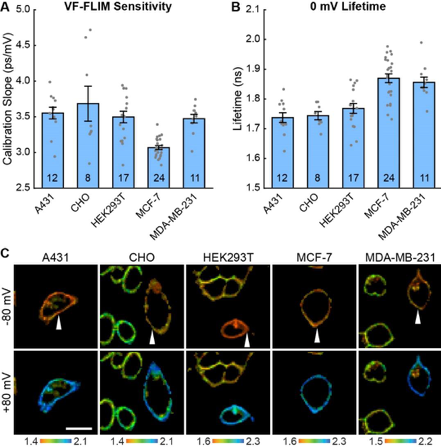
Figure 1. VF-FLIM is a general and portable method for optically determining membrane potential. VF2.1.Cl lifetime-voltage relationships were determined with whole cell voltage clamp electrophysiology in five cell lines. (A) Slope and (B) 0 mV reference point of linear fits for the lifetime-voltage relationship, shown as mean ± S.E.M. Gray dots are single cells. (C) Representative lifetime-intensity overlay images for each cell line with the indicated cells (white arrow) held at -80 mV (top) or +80 mV (bottom). Lifetime scales are in ns. Scale bar is 20 μm. Reproduced from Figure 2. of the preprint
Membrane potential dynamics in epidermal growth factor signaling
The authors next used VF-FILM to elucidate the role of Vmemduring EGF/EGF receptor (EGFR)-mediated signaling in A431 cells. EGF treatment results in a 15mV hyperpolarization within 60-90 seconds in approximately 80% of cells followed by a return to baseline in 15 minutes (Fig 2). Inhibition of EGFR and ErbB2 tyrosine kinase activity with the covalent inhibitor canertinib abolishes hyperpolarization. Additionally, blockade of the EGFR kinase domain with gefitnib also diminishes hyperpolarization indicating that A431 cells exhibit an EGF-induced hyperpolarization which is dependent on the kinase activity of EGFR.
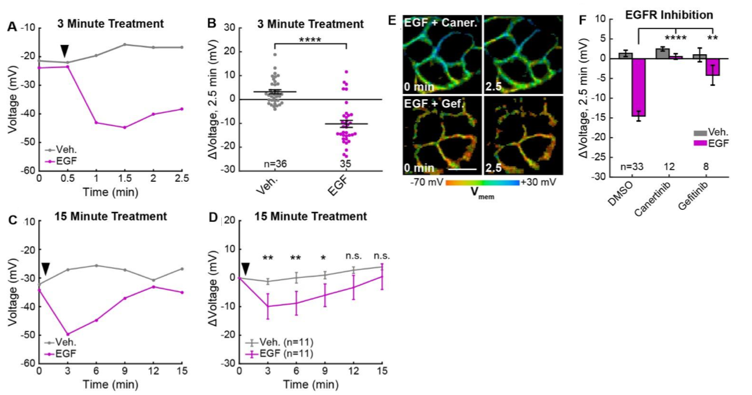
Figure. 2 EGFR-mediated receptor tyrosine kinase activity produces a transient hyperpolarization in A431 cells. (A) Quantification of images. Vehicle (Veh.)/EGF added at black arrow. (B) Aggregated responses for various trials of cells treated with vehicle or EGF. (C) Quantification of images with Veh/EGF. (D) Average response of cells. (E) Lifetime images of A431 cells before and after EGF addition, with 500 nM canertinib (top) or 10 μM gefitinib (bottom). (F) Voltage changes 2.5 minutes after EGF addition in cells treated with DMSO (vehicle control) or an EGFR inhibitor. Scale bar is 20 μm. (C,F,H): Asterisks indicate significant differences between vehicle and EGF at that time point. (F): Asterisks reflect significant differences between EGF-induced voltage responses with DMSO vehicle or an EGFR inhibitor (n.s. p>0.05, * p<0.05, ** p<0.01, *** p<0.001, **** p<0.0001, two-tailed, unpaired, unequal variances t-test). Reproduced from Figure 4. of the preprint
The authors identified the outward potassium channels as the mediating current of EGF-induced hyperpolarization. Block of K+ driving force with high extracellular [K+] completely abolished the EGF-induced hyperpolarization response whereas blockage of voltage-gated K+channels with 4-aminopyridine (4-AP) enhances the hyperpolarization. Specific inhibition of intermediate-conductance Ca2+-activated potassium channel KCa3.1 using TRAM-43 also abolished EGF-induced hyperpolarization. The hyperpolarizing current was found to be carried by K+ions passed through Ca2+-activated K+channel KCa3.1 and mediated by intracellular Ca2+stores. In the context of receptor tyrosine kinase signalling, Vmem may modulate the driving force for external Ca2+entry and act as a regulator for Ca2+signalling. Therefore, VF-FLIM could be a useful tool in assessing effects of small Vmemchanges on signaling pathways in non-excitable cells.
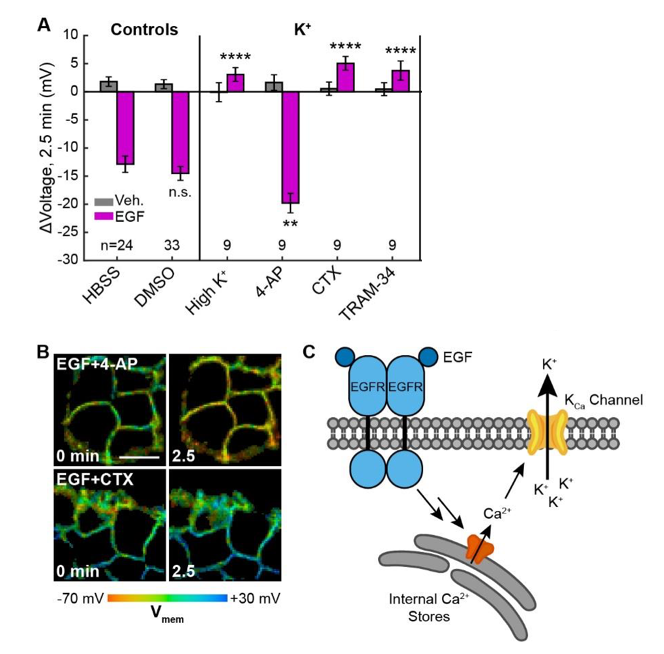
Figure 3.EGF-induced hyperpolarization is mediated by a Ca activated K channel. (A) Comparison of the Vmem change 2.5 minutes after EGF addition in cells incubated in unmodified imaging buffer (HBSS) or in modified solutions. (B) Lifetime images of A431 cells treated with 4-AP or CTX. (C) Model for membrane hyperpolarization following EGFR activation. Scale bar is 20 μm. Bars are mean ± SEM. Asterisks reflect significant differences in EGF- stimulated Vmem change between the unmodified control (HBSS or DMSO) and modified solutions (n.s. p>0.05, * p<0.05, ** p<0.01, *** p<0.001, **** p<0.0001, two-tailed, unpaired, unequal variances t-test). Reproduced from Figure 5. of the preprint
Why I liked this preprint
This is a nice article that shows in a simple way that membrane potential can be determined using VF-FLIM but that it can also be used to probe membrane potential in cellular states that are not accessible or challenging with standard electrophysiological techniques. The authors make a real effort to validate their method through several experiments and provide plenty of figures in the supplementary data to get a real understanding of the probe and its uses.
Questions to authors
1) Cells in culture or in primary tissue are electrically connected via gap junctions, however VF-FLIM was found to be robust in small groups of cells. How many cells can be used in a population type recording and what are the spatial limitations of the technique?
2) In a long-term experiment how do you control for probe stability over time and exclude ion channel run down?
3) Is there a temperature dependency of the fluorescence of VF-FLIM?
References
- Ouadid-ahidouch, H., Bourhis, X. Le & Roudbaraki, M. Changes in the K + current-density of MCF-7 cells during progression through the cell cycle : Possible Involvement of a h-ether . a-gogo K + channel. (2001).
- Byrd, R. C. & Sciences, H. Changes in Membrane Potential During the Progression of MCF-7 Human Mammary Tumor Cells Through the Cell Cycle. 185,177–185 (1995).
- Flag, T., Oh, L.-, Gst, T. & Web, W. Letters To Nature. Nature408,381–386 (2000).
- Horn, R. & Korn, S. J. Prevention of rundown in electrophysiological recording. Methods Enzymol.207,149–155 (1992).
doi: https://doi.org/10.1242/prelights.9406
Read preprintSign up to customise the site to your preferences and to receive alerts
Register hereAlso in the bioengineering category:
Detergent-Triggered Membrane Remodelling Monitored via Intramembrane Fluorescence De-Quenching
Cyntia Alves Conceição, Marcus Oliveira
A Novel Chimeric Antigen Receptor (CAR) - Strategy to Target EGFRVIII-Mutated Glioblastoma Cells via Macrophages
Dina Kabbara
Human pluripotent stem cell-derived macrophages modify development of human kidney organoids
Theodora Stougiannou
Also in the molecular biology category:
Remote homology and functional genetics unmask deeply preserved Scm3/HJURP orthologs in metazoans
Reinier Prosee
Exogenous Amyloid Sequences: Their Role in Amyloid-Beta Heterotypic Aggregation
Cyntia Alves Conceição, Marcus Oliveira
Detergent-Triggered Membrane Remodelling Monitored via Intramembrane Fluorescence De-Quenching
Cyntia Alves Conceição, Marcus Oliveira
Also in the physiology category:
Resilience to cardiac aging in Greenland shark Somniosus microcephalus
Theodora Stougiannou
Trade-offs between surviving and thriving: A careful balance of physiological limitations and reproductive effort under thermal stress
Tshepiso Majelantle
Imaging cellular activity simultaneously across all organs of a vertebrate reveals body-wide circuits
Muhammed Sinan Malik
preLists in the bioengineering category:
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Advances in microscopy
This preList highlights exciting unpublished preprint articles describing advances in microscopy with a focus on light-sheet microscopy.
| List by | Stephan Daetwyler |
Also in the molecular biology category:
BSDB Spring Meeting: Molecules to Morphogenesis
The British Society for Developmental Biology (BSDB) Spring Meeting Molecules to Morphogenesis was held from 23–26 March 2026 at the University of Warwick (UK). This meeting brought together a vibrant community of researchers to discuss how molecular mechanisms are integrated across scales to drive morphogenesis, spanning diverse model systems and approaches. This preList contains preprints by presenters from the talk and poster sessions at the meeting. Please do get in touch at preLights@biologists.com if you notice any relevant preprints that we may have missed.
| List by | Ingrid Tsang |
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
2024 Hypothalamus GRC
This 2024 Hypothalamus GRC (Gordon Research Conference) preList offers an overview of cutting-edge research focused on the hypothalamus, a critical brain region involved in regulating homeostasis, behavior, and neuroendocrine functions. The studies included cover a range of topics, including neural circuits, molecular mechanisms, and the role of the hypothalamus in health and disease. This collection highlights some of the latest advances in understanding hypothalamic function, with potential implications for treating disorders such as obesity, stress, and metabolic diseases.
| List by | Nathalie Krauth |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |
Also in the physiology category:
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Cellular metabolism
A curated list of preprints related to cellular metabolism at Biorxiv by Pablo Ranea Robles from the Prelights community. Special interest on lipid metabolism, peroxisomes and mitochondria.
| List by | Pablo Ranea Robles |











 (3 votes)
(3 votes)