Oxidative stress triggers RNAPII arrest through PARylation and DNA damage
Posted on: 1 April 2026
Preprint posted on 6 November 2025
Attachment of ADP-ribose molecules to RNA Polymerase II allows its traffic regulation on transcribed genes in response to oxidative DNA damage
Selected by Élise Jeanne Pouponnot, Pierre CaronCategories: cell biology
Background
Every day, our cells experience tens of thousands of DNA lesions (1). These include base modifications arising from oxidative stress or from exposure to exogenous agents such as certain anticancer therapies. To counteract these small and non-helix-distorting DNA lesions, cells rely on the Base Excision Repair (BER) pathway (2).
This pathway begins when specialized enzymes called DNA glycosylases (such as 8-Oxoguanine DNA Glycosylase, OGG1) scan DNA for damaged bases caused by oxidation, such as 8-oxodG. Once detected, these enzymes remove the damaged base, leaving behind a missing-base site known as an abasic (AP) site.
These abasic sites are then cut by either DNA glycosylases with additional “cutting” activity or by the enzyme apurinic/apyrimidinic endonuclease 1 (APE1), which produces a small break in one strand of the DNA (2). This break serves as a signal to recruit poly(ADP-ribose) polymerase 1 (PARP1). PARP1 detects the damage and modifies itself and nearby histones with ADP-ribose chains (e.g. PARylation), causing the surrounding chromatin to loosen. This relaxed chromatin environment allows other repair proteins to access the site and complete DNA repair through DNA synthesis and ligation (3).
Approximately 2% of the human genome consists of genes that can be transcribed by the RNA polymerase II complex (RNAPII). When transcribed genes are damaged in their body by DNA double-strand breaks (DSBs) or UV-induced DNA lesions, RNAPII progression is blocked at damage sites and mRNA production is consequently interrupted (4, 5).
Interestingly, whereas some types of base oxidation do not impede RNAPII progression, others do just like repair intermediates of oxidized bases, e.g. AP sites and ssDNA breaks. Although transcription-coupled BER has been the subject of several studies (6, 7), the factors involved, and the regulatory mechanisms remain largely unknown.
In this preprint (8), Thomas and colleagues investigate how oxidative stress affects transcription by mapping RNAPII and nascent transcription in a genome-wide manner. The authors show that active RNAPII is transiently stalled at defined positions within oxidatively damaged genes, with promoter-proximal RNAPII pausing dynamically repositioned in a NELF-dependent manner. Importantly, transcriptional restart requires the removal of PARylation chains by PARG and the efficient repair of oxidative lesions by the BER factors OGG1 and X-Ray Repair Cross Complementing 1 (XRCC1). Together, these findings provide a refined and spatially resolved view of how transcription is dynamically coordinated with BER to enable rapid recovery of gene expression following oxidative stress.
Key findings
- RNAPII arrested within gene bodies can locally resume transcription without new initiation
To determine whether transcriptional recovery requires new RNAPII initiation, the authors combined pharmacological inhibition of promoter escape using the CDK9 inhibitor DRB with genome-wide nascent transcription profiling (TT-seq). By blocking the release of newly initiated RNAPII complexes from promoters, they show that transcriptional recovery still occurred following oxidative stress, demonstrating that RNAPII complexes arrested by oxidative damage can restart transcription directly from within gene bodies. Moreover, TT-seq profiles during recovery revealed a pronounced bias of nascent transcription toward downstream regions of genes, further supporting a local restart mechanism.
Together, these results uncover a mode of transcriptional recovery that bypasses promoter-dependent initiation and enables rapid restoration of gene expression.
- Promoter-proximal regulation of early RNAPII elongation by NELF and PARylation
To dissect the mechanisms underlying transcriptional repression and recovery following oxidative stress, the authors combined genome-wide profiling of RNAPII occupancy (ECLAP-seq) with nascent transcription analysis (TT-seq). At the promoter-proximal region, analysis of RNAPII distribution around transcription start sites (TTS) revealed that oxidative stress induces a redistribution of paused RNAPII complexes (Figure 1A). By depleting the pausing factor NELF, the authors show that NELF is not required for the global shutdown of transcription. Instead, it modulates the positioning of early elongation complexes, promoting a transient shift of RNAPII pausing from the canonical first pause site to a second pause site associated with the +1 nucleosome during stress and recovery. In addition, using pharmacological inhibition of PARP1, the authors demonstrate that PARylation acts as a reversible brake on early RNAPII elongation. High levels of PARylation coincide with transcriptional repression and blocking PARP1 activity promotes the release of RNAPII complexes, without preventing the initial transcriptional shutdown. Finally, transcriptional recovery requires the active removal of PAR chains by the poly(ADP-ribose) glycohydrolase PARG, indicating that PARylation must be rapidly reversed to permit RNAPII reactivation (Figure 1B).
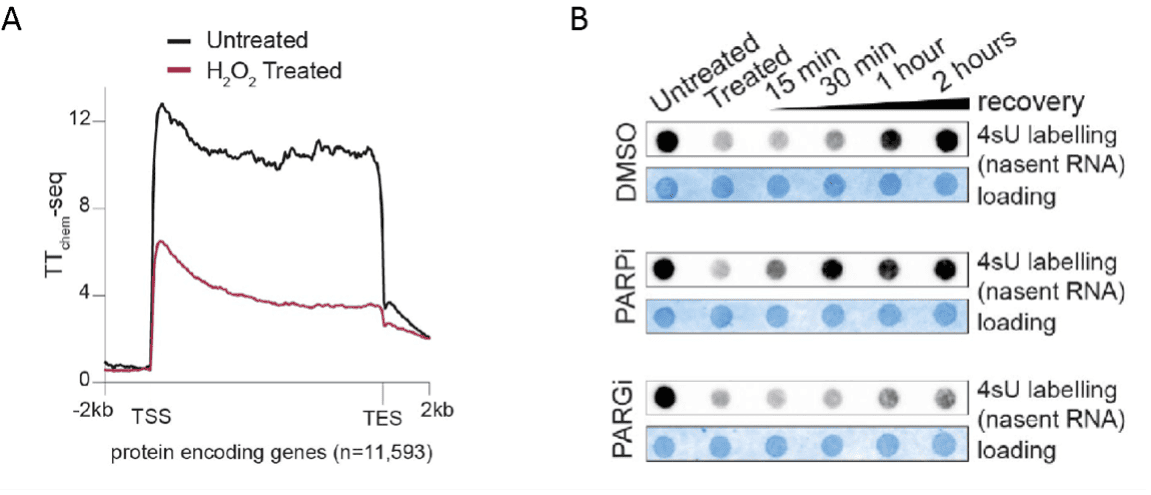
Figure 1. (from fig. 1.g and fig. 5.c, Thomas et al. 2025).
(A) Metagene profiles of nascent transcription measured by TT-chem-seq across protein-coding genes in untreated cells (dark line) or following H₂O₂-induced oxidative stress (red line). Oxidative damage induces a strong and global reduction of RNA synthesis along gene bodies while preserving RNAPII occupancy at transcription start sites (TSS), consistent with transcriptional arrest without RNAPII dissociation.
(B) Dot blot analysis of 4-thiouridine (4sU)–labelled nascent RNA in control (DMSO), PARP-inhibited (PARPi) or PARG-inhibited (PARGi) cells following oxidative stress and during recovery. In control cells, nascent transcription is transiently reduced after stress and progressively recovers over time. PARP inhibition accelerates this recovery, whereas inhibition of PARG strongly impairs transcriptional restart. These results indicate that dynamic PARylation, and in particular the timely removal of PAR chains, is required for efficient transcriptional recovery. Methylene blue staining is shown as a loading control.
- DNA repair–dependent restart of RNAPII within gene bodies
Remarkably, the authors also unveil that the recovery of transcription depends on DNA repair. Using XRCC1-deficient cells and pharmacological inhibition of the DNA glycosylase OGG1, the authors show that efficient repair of oxidative DNA lesions is required to relieve RNAPII blocks within gene bodies. Their activity enables RNAPII to locally resume transcription from within genes, independently of new transcription initiation.
Together, the findings described in sections 2 and 3 show that transcriptional recovery relies on two coordinated layers of regulation: PARylation-dependent control of early RNAPII elongation near promoters, and DNA repair–dependent removal of physical blocks that enables RNAPII to restart within gene bodies.
What I like about this preprint
By combining methods that measure nascent RNA synthesis with approaches that map RNAPII occupancy, the authors can distinguish changes in transcriptional activity from changes in polymerase binding to chromatin. This strategy provides a clear view of how RNAPII behavior is dynamically regulated after oxidative stress and helps link transcriptional regulation with DNA repair in a spatially resolved manner.
I really liked this preprint for several reasons, such as the fact that it elevates BER/SSBR from “genome maintenance” to active regulators of transcriptional dynamics and it shows that “global repression” doesn’t mean “global chaos.” It’s structured, modular, and reversible. What really got me excited isn’t just the data, it is that the story feels right, it explains a big, common biological phenomenon (oxidative stress) without invoking exotic machinery.
Question for the authors
Q1: What do you think is the direct PARylation target responsible for restricting early RNAPII release?
Q2: Do you think PARP2 could also be involved in the RNAPII positioning on damaged genes?
References
- LINDAHL,T. and BARNES,D.E. (2000) Repair of Endogenous DNA Damage. Cold Spring Harb Symp Quant Biol, 65, 127–134.
- Krokan,H.E. and Bjoras,M. (2013) Base Excision Repair. Cold Spring Harb Perspect Biol, 5, a012583–a012583.
- Özdemir,C., Purkey,L.R., Sanchez,A. and Miller,K.M. (2024) PARticular MARks: Histone ADP-ribosylation and the DNA damage response. DNA Repair (Amst), 140, 103711.
- Lesage,E., Clouaire,T. and Legube,G. (2021) Repair of DNA double-strand breaks in RNAPI- and RNAPII-transcribed loci. DNA Repair (Amst), 104, 103139.
- van der Meer,P.J. and Luijsterburg,M.S. (2025) The molecular basis of human transcription-coupled DNA repair. Nat Cell Biol, 10.1038/s41556-025-01715-9.
- Bilkis,R., Lake,R.J., Cooper,K.L., Tomkinson,A. and Fan,H.-Y. (2023) The CSB chromatin remodeler regulates PARP1- and PARP2-mediated single-strand break repair at actively transcribed DNA regions. Nucleic Acids Res, 51, 7342–7356.
- Heine,G.F., Horwitz,A.A. and Parvin,J.D. (2008) Multiple Mechanisms Contribute to Inhibit Transcription in Response to DNA Damage. Journal of Biological Chemistry, 283, 9555–9561.
- Thomas,Q.A., Wu,L., Lesage,E., Iversen,H.K.M., López Martínez,D., Kompocholi,S., Liu,H., Nieto Moreno,N. and Gregersen,L.H. (2025) Oxidative stress triggers RNAPII arrest through PARylation and DNA damage. 10.1101/2025.11.06.686939.
Sign up to customise the site to your preferences and to receive alerts
Register hereAlso in the cell biology category:
Classical enhancers couple cis-regulatory logic with transcriptional condensates and 3D genome architecture
Siddharth Singh
Small Molecule Agonists of TREM2 Reprogram Microglia and Protect Synapses in Human Alzheimer’s Models
Dina Kabbara
Cell-intrinsic compliance mechanism enables release of tensile stress to prevent tissue rupture
Ruoheng Li
preLists in the cell biology category:
Keystone Symposium on Stem Cell Models in Embryology 2026
The Keystone Symposium on Stem Cell Models in Embryology, 2026, was organised by Jun Wu (UT Southwestern), Jianping Fu (University of Michigan) and Miki Ebisuya (TU Dresden) and held at Asilomar Conference Grounds in California (US). The meeting discussed recent advances made in establishing stem-cell-based embryo models, what fundamental insights into developmental processes have been gleaned from them, as well as how they are beginning to be applied more widely. This prelist contains preprints by presenters at the talk and poster sessions at the conference, which our Reviews Editor in attendance spotted. Please do reach out to preLights@biologists.com if you notice any that we’ve missed.
| List by | Ingrid Tsang |
SciELO preprints – From 2025 onwards
SciELO has become a cornerstone of open, multilingual scholarly communication across Latin America. Its preprint server, SciELO preprints, is expanding the global reach of preprinted research from the region (for more information, see our interview with Carolina Tanigushi). This preList brings together biological, English language SciELO preprints to help readers discover emerging work from the Global South. By highlighting these preprints in one place, we aim to support visibility, encourage early feedback, and showcase the vibrant research communities contributing to SciELO’s open science ecosystem.
| List by | Carolina Tanigushi |
November in preprints – DevBio & Stem cell biology
preLighters with expertise across developmental and stem cell biology have nominated a few developmental and stem cell biology (and related) preprints posted in November they’re excited about and explain in a single paragraph why. Concise preprint highlights, prepared by the preLighter community – a quick way to spot upcoming trends, new methods and fresh ideas.
| List by | Aline Grata et al. |
October in preprints – DevBio & Stem cell biology
Each month, preLighters with expertise across developmental and stem cell biology nominate a few recent developmental and stem cell biology (and related) preprints they’re excited about and explain in a single paragraph why. Short, snappy picks from working scientists — a quick way to spot fresh ideas, bold methods and papers worth reading in full. These preprints can all be found in the October preprint list published on the Node.
| List by | Deevitha Balasubramanian et al. |
October in preprints – Cell biology edition
Different preLighters, with expertise across cell biology, have worked together to create this preprint reading list for researchers with an interest in cell biology. This month, most picks fall under (1) Cell organelles and organisation, followed by (2) Mechanosignaling and mechanotransduction, (3) Cell cycle and division and (4) Cell migration
| List by | Matthew Davies et al. |
September in preprints – Cell biology edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading list. This month, categories include: (1) Cell organelles and organisation, (2) Cell signalling and mechanosensing, (3) Cell metabolism, (4) Cell cycle and division, (5) Cell migration
| List by | Sristilekha Nath et al. |
July in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell Signalling and Mechanosensing (2) Cell Cycle and Division (3) Cell Migration and Cytoskeleton (4) Cancer Biology (5) Cell Organelles and Organisation
| List by | Girish Kale et al. |
June in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: (1) Cell organelles and organisation (2) Cell signaling and mechanosensation (3) Genetics/gene expression (4) Biochemistry (5) Cytoskeleton
| List by | Barbora Knotkova et al. |
May in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) Biochemistry/metabolism 2) Cancer cell Biology 3) Cell adhesion, migration and cytoskeleton 4) Cell organelles and organisation 5) Cell signalling and 6) Genetics
| List by | Barbora Knotkova et al. |
Keystone Symposium – Metabolic and Nutritional Control of Development and Cell Fate
This preList contains preprints discussed during the Metabolic and Nutritional Control of Development and Cell Fate Keystone Symposia. This conference was organized by Lydia Finley and Ralph J. DeBerardinis and held in the Wylie Center and Tupper Manor at Endicott College, Beverly, MA, United States from May 7th to 9th 2025. This meeting marked the first in-person gathering of leading researchers exploring how metabolism influences development, including processes like cell fate, tissue patterning, and organ function, through nutrient availability and metabolic regulation. By integrating modern metabolic tools with genetic and epidemiological insights across model organisms, this event highlighted key mechanisms and identified open questions to advance the emerging field of developmental metabolism.
| List by | Virginia Savy, Martin Estermann |
April in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell cycle and division 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) (epi)genetics
| List by | Vibha SINGH et al. |
March in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) cancer biology 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics and genomics 6) other
| List by | Girish Kale et al. |
Biologists @ 100 conference preList
This preList aims to capture all preprints being discussed at the Biologists @100 conference in Liverpool, UK, either as part of the poster sessions or the (flash/short/full-length) talks.
| List by | Reinier Prosee, Jonathan Townson |
February in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry and cell metabolism 2) cell organelles and organisation 3) cell signalling, migration and mechanosensing
| List by | Barbora Knotkova et al. |
Community-driven preList – Immunology
In this community-driven preList, a group of preLighters, with expertise in different areas of immunology have worked together to create this preprint reading list.
| List by | Felipe Del Valle Batalla et al. |
January in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) biochemistry/metabolism 2) cell migration 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Barbora Knotkova et al. |
December in preprints – the CellBio edition
A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. This month, categories include: 1) cell cycle and division 2) cell migration and cytoskeleton 3) cell organelles and organisation 4) cell signalling and mechanosensing 5) genetics/gene expression
| List by | Matthew Davies et al. |
November in preprints – the CellBio edition
This is the first community-driven preList! A group of preLighters, with expertise in different areas of cell biology, have worked together to create this preprint reading lists for researchers with an interest in cell biology. Categories include: 1) cancer cell biology 2) cell cycle and division 3) cell migration and cytoskeleton 4) cell organelles and organisation 5) cell signalling and mechanosensing 6) genetics/gene expression
| List by | Felipe Del Valle Batalla et al. |
BSCB-Biochemical Society 2024 Cell Migration meeting
This preList features preprints that were discussed and presented during the BSCB-Biochemical Society 2024 Cell Migration meeting in Birmingham, UK in April 2024. Kindly put together by Sara Morais da Silva, Reviews Editor at Journal of Cell Science.
| List by | Reinier Prosee |
‘In preprints’ from Development 2022-2023
A list of the preprints featured in Development's 'In preprints' articles between 2022-2023
| List by | Alex Eve, Katherine Brown |
preLights peer support – preprints of interest
This is a preprint repository to organise the preprints and preLights covered through the 'preLights peer support' initiative.
| List by | preLights peer support |
The Society for Developmental Biology 82nd Annual Meeting
This preList is made up of the preprints discussed during the Society for Developmental Biology 82nd Annual Meeting that took place in Chicago in July 2023.
| List by | Joyce Yu, Katherine Brown |
CSHL 87th Symposium: Stem Cells
Preprints mentioned by speakers at the #CSHLsymp23
| List by | Alex Eve |
Journal of Cell Science meeting ‘Imaging Cell Dynamics’
This preList highlights the preprints discussed at the JCS meeting 'Imaging Cell Dynamics'. The meeting was held from 14 - 17 May 2023 in Lisbon, Portugal and was organised by Erika Holzbaur, Jennifer Lippincott-Schwartz, Rob Parton and Michael Way.
| List by | Helen Zenner |
9th International Symposium on the Biology of Vertebrate Sex Determination
This preList contains preprints discussed during the 9th International Symposium on the Biology of Vertebrate Sex Determination. This conference was held in Kona, Hawaii from April 17th to 21st 2023.
| List by | Martin Estermann |
Alumni picks – preLights 5th Birthday
This preList contains preprints that were picked and highlighted by preLights Alumni - an initiative that was set up to mark preLights 5th birthday. More entries will follow throughout February and March 2023.
| List by | Sergio Menchero et al. |
CellBio 2022 – An ASCB/EMBO Meeting
This preLists features preprints that were discussed and presented during the CellBio 2022 meeting in Washington, DC in December 2022.
| List by | Nadja Hümpfer et al. |
Fibroblasts
The advances in fibroblast biology preList explores the recent discoveries and preprints of the fibroblast world. Get ready to immerse yourself with this list created for fibroblasts aficionados and lovers, and beyond. Here, my goal is to include preprints of fibroblast biology, heterogeneity, fate, extracellular matrix, behavior, topography, single-cell atlases, spatial transcriptomics, and their matrix!
| List by | Osvaldo Contreras |
EMBL Synthetic Morphogenesis: From Gene Circuits to Tissue Architecture (2021)
A list of preprints mentioned at the #EESmorphoG virtual meeting in 2021.
| List by | Alex Eve |
FENS 2020
A collection of preprints presented during the virtual meeting of the Federation of European Neuroscience Societies (FENS) in 2020
| List by | Ana Dorrego-Rivas |
Planar Cell Polarity – PCP
This preList contains preprints about the latest findings on Planar Cell Polarity (PCP) in various model organisms at the molecular, cellular and tissue levels.
| List by | Ana Dorrego-Rivas |
BioMalPar XVI: Biology and Pathology of the Malaria Parasite
[under construction] Preprints presented at the (fully virtual) EMBL BioMalPar XVI, 17-18 May 2020 #emblmalaria
| List by | Dey Lab, Samantha Seah |
1
Cell Polarity
Recent research from the field of cell polarity is summarized in this list of preprints. It comprises of studies focusing on various forms of cell polarity ranging from epithelial polarity, planar cell polarity to front-to-rear polarity.
| List by | Yamini Ravichandran |
TAGC 2020
Preprints recently presented at the virtual Allied Genetics Conference, April 22-26, 2020. #TAGC20
| List by | Maiko Kitaoka et al. |
3D Gastruloids
A curated list of preprints related to Gastruloids (in vitro models of early development obtained by 3D aggregation of embryonic cells). Updated until July 2021.
| List by | Paul Gerald L. Sanchez and Stefano Vianello |
ECFG15 – Fungal biology
Preprints presented at 15th European Conference on Fungal Genetics 17-20 February 2020 Rome
| List by | Hiral Shah |
ASCB EMBO Annual Meeting 2019
A collection of preprints presented at the 2019 ASCB EMBO Meeting in Washington, DC (December 7-11)
| List by | Madhuja Samaddar et al. |
EMBL Seeing is Believing – Imaging the Molecular Processes of Life
Preprints discussed at the 2019 edition of Seeing is Believing, at EMBL Heidelberg from the 9th-12th October 2019
| List by | Dey Lab |
Autophagy
Preprints on autophagy and lysosomal degradation and its role in neurodegeneration and disease. Includes molecular mechanisms, upstream signalling and regulation as well as studies on pharmaceutical interventions to upregulate the process.
| List by | Sandra Malmgren Hill |
Lung Disease and Regeneration
This preprint list compiles highlights from the field of lung biology.
| List by | Rob Hynds |
Cellular metabolism
A curated list of preprints related to cellular metabolism at Biorxiv by Pablo Ranea Robles from the Prelights community. Special interest on lipid metabolism, peroxisomes and mitochondria.
| List by | Pablo Ranea Robles |
BSCB/BSDB Annual Meeting 2019
Preprints presented at the BSCB/BSDB Annual Meeting 2019
| List by | Dey Lab |
MitoList
This list of preprints is focused on work expanding our knowledge on mitochondria in any organism, tissue or cell type, from the normal biology to the pathology.
| List by | Sandra Franco Iborra |
Biophysical Society Annual Meeting 2019
Few of the preprints that were discussed in the recent BPS annual meeting at Baltimore, USA
| List by | Joseph Jose Thottacherry |
ASCB/EMBO Annual Meeting 2018
This list relates to preprints that were discussed at the recent ASCB conference.
| List by | Dey Lab, Amanda Haage |











 (No Ratings Yet)
(No Ratings Yet)